Water supply may influence the bacterioplankton community structure

The use of high fish densities during production may result in more rapid and severe disease outbreaks. Outbreaks of parasitic, bacterial and fungal diseases are among the most important limiting factors for the success of aquaculture production, leading to high mortality rates and important economic losses. For example, the production of Senegalese sole (Solea senegalensis), a species of considerable commercial value, is strongly limited by its sensitivity to infectious diseases such as pasteurellosis, vibriosis and flexibacteriosis.
The aquaculture water microbiome is essential for maintaining water quality (nutrient recycling) and fish health during intensive fish production. Naturally occurring or introduced beneficial bacteria (probiotics) may contribute to improve water quality, inhibit the development of fish pathogens, improve the fish immune system and promote the balance of the fish bacterial flora.
In previous studies, we showed that S. senegalensis appears to influence the bacterial communities in a grow-out RAS and that, despite the presence of several potential fish pathogens, no diseased fish were observed during the study period. Our findings indicated that the water in grow-out RAS was dominated by naturally occurring beneficial microbes (antagonistic populations), which may have played an important role in suppressing the development of putative pathogens. However, we could not determine if such a trend would also be detectable in RAS systems used for production of juvenile specimens (hatchery), which are managed under different conditions and stocked from hatching until juvenile stage.
This article – adapted and summarized from the original – reports on an investigation of bacterioplankton community composition and diversity in the water of a commercial hatchery operating a RAS for the production of Senegalese sole juveniles, and compares results with those previously recorded for sole grow-out in RAS. We also evaluated our results in light of the putative function of bacterioplankton populations in the hatchery RAS.
Study setup
Fieldwork was conducted at a RAS hatchery using shallow raceway systems (SRS) for production of more than 1 million Senegalese sole juveniles annually, grown from hatching to a weight of about 40 grams. The hatchery recirculated water at less than 5 percent of total system volume per day.
The water supply reservoir (Sup) is filled with seawater pumped through an inlet pipe from the ocean and ozonized in a tank connected to a protein skimmer (Ozo) before entering the pre-production reservoir (Pre), the hatchery with ~4-gram juvenile sole at densities about 3.7 kg per square meter. Water from Pre is recycled by passing through a sedimentation tank (Sed) where mechanical filtration is also carried out. After mechanical filtration, water flows to a biofilter tank (Bio) for biological filtration and is subsequently pumped back to Ozo where it reenters the system. Water samples for bacterial community analysis and chemical characterization were collected in triplicate from all five different compartments (Sup, Ozo, Pre, Sed and Bio), and several analyzes performed.
For additional information on the study setup, including water chemistry analysis, DNA analysis, and bacterial communities, please refer to the original publication.
Results and discussion
DGGE analysis (PCR technique used to identify microbial populations) of bacterioplankton communities showed that, despite the young age of fishes and their relatively short period in the tanks (45 days), there was a significant separation between water supply and RAS compartments. The communities of RAS compartments defined by DGGE also tended to cluster together.
To examine changes in bacterial richness, rarefaction curves (technique to assess species richness for a given number of individual samples) were generated for all compartments (Fig. 1). Controlling for sampling size, operational taxonomic unit (OTU; an operational definition used to classify groups of closely related individuals) richness in the Sup compartment was 35.79 ± 1.02. In the aquaculture tanks, richness was lowest in Sed (69.84 ± 1.74) and highest in Ozo (92.88 ± 4.82). The high diversity detected in Ozo may be due to an important fraction of dead microorganisms that accumulate in this compartment naturally derived from water supply and fish and feed waste from Pre tank and from the bacteria that proliferate in the biofilter.
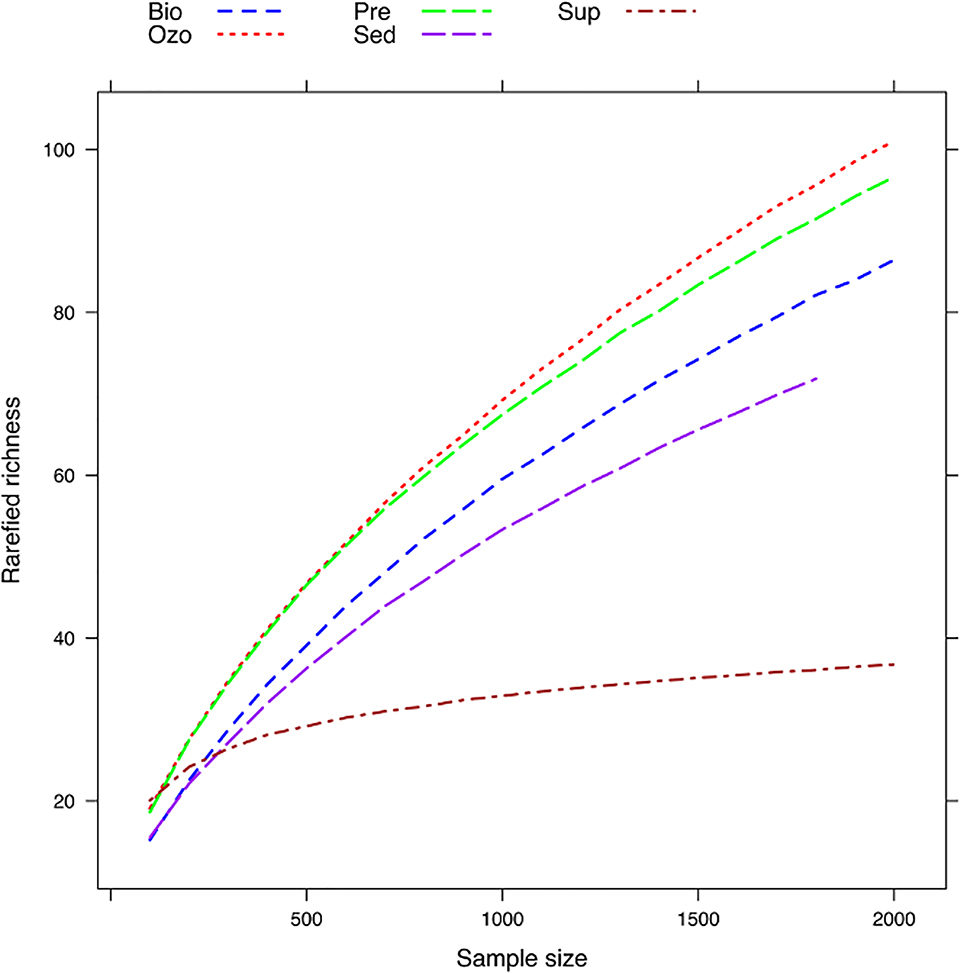
Results indicate that, despite the fact that the bacterioplankton communities in the water supply were clearly distinct from RAS tanks, several dominant bacterial communities in the hatchery tanks were originally introduced in the system through the water supply. This finding is in contrast with the results obtained for sole grow-out RAS, where only few bacterial OTUs were found to be dominant in the water supply and fish tanks. Probably, due to the early life stage development of the fish in this study, gut microbes released to the environment via feces may have had lower influence on hatchery water bacterioplankton than in grow-out RAS.
Fig. 2 shows taxonomic information about the most abundant OTUs and phylogenetic analyses to identify ecotypes (genetically distinct geographic varieties, populations or races within a species, genotypically adapted to specific environmental conditions) related to these OTUs in different RAS compartments. This approach allowed us to better understand the composition and putative ecological role of the dominant bacterial populations in the RAS bacterioplankton.
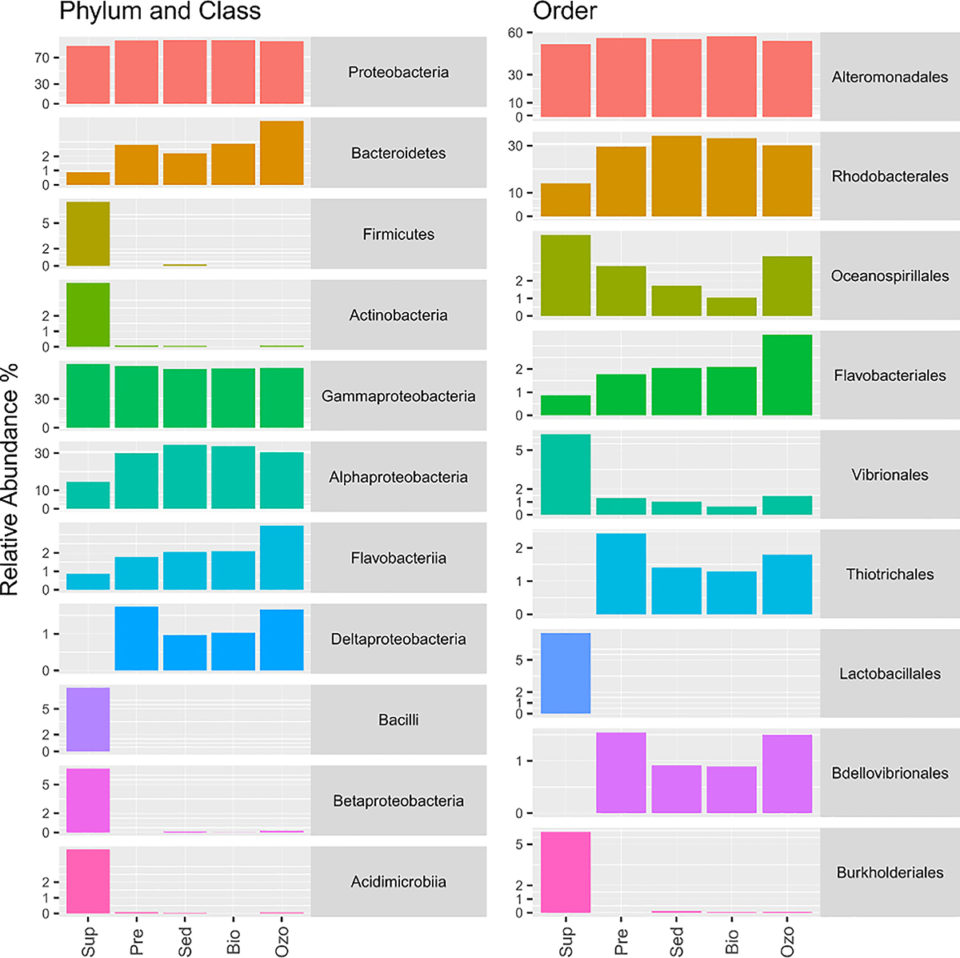
The overall taxonomic analyses showed that Proteobacteria was the most abundant bacterial phylum in all RAS compartments (average relative abundance 94.60 ± 4.10 percent), followed by Bacteroidetes (average relative abundance 2.65 ±1.30 percent). The phyla Firmicutes and Actinobacteria were more abundant in the water supply (7.50 percent and 4.13 percent, respectively) than in the hatchery RAS (0.04 percent and 0.05 percent, respectively). The most abundant orders detected in this study were Alteromonadales (54.98 ± 2.16 percent), Rhodobacterales (28.22 ± 8.17 percent), Oceanospirillales (2.73 ± 1.41 percent), Vibrionales (2.14 ± 2.32 percent), Flavobacteriales (2.05 ± 0.94 percent), Lactobacillales (1.50 ± 3.35 percent), Thiotrichales (1.38 ± 0.89 percent), Burkholderiales (1.22 ± 2.62 percent) and Bdellovibrionales (0.97±0.62 percent).
Overall, our results showed that, with exception of two, all dominant OTUs detected in the hatchery tanks were originally present in the water supply before entering the RAS. To evaluate the composition of potential fish pathogens in the hatchery RAS we also specifically searched for OTUs related to bacterial genera which often comprise known fish pathogens. We identified Vibrio ichthyoenteri, previously reported to be a pathogen of olive flounder.
Likewise, we identified a microorganism similar to Vibrio anguillarum, a pathogen that causes vibriosis in approximately 50 species of fish. Two other pathogens, Serratia marcescens and Francisella philomiragia, were identified in low abundance inside the RAS and only the former was detected in the fish compartment (Pre). This species is an opportunistic waterborne pathogen able to cause disease in a range of animals, including finfish species. However, in line with our previous study and despite the presence of potential pathogens, no disease outbreak occurred in the hatchery RAS during this study.
Perspectives
Exploring the potential of naturally occurring microorganisms as biocontrol agents in aquacultures is not a new concept. The development of microbial management or modulation approaches should be based on a fundamental knowledge about the aquaculture microbiome.
This study provides baseline information about the bacterioplankton community composition and diversity of a commercial hatchery RAS for the production of juvenile Senegalese sole. Our results showed that despite the differences in relative abundance, the most abundant orders detected in the hatchery RAS (Alteromonadales, Rhodobacterales, Oceanospirillales, Vibrionales and Flavobacteriales) were also the most abundant detected in the sole grow-out RAS characterized in our previous study.
Curiously, in contrast to our findings for grow-out RAS, our results indicated that the bacterial assemblage of the water supply played an important role for the colonization of bacterial populations in the hatchery RAS. Most remarkable, here water supply seems to contribute for a strong colonization of Pseudoalteromonas sp. in the tanks, which in turn may play a role in suppressing the development of potential fish pathogens in the aquaculture system.
Our results suggest that the bacterial community of the water supply may influence the bacterioplankton community structure of sole hatchery RAS. However, considering the results obtained for sole grow-out RAS, the contribution of the water supply to shape RAS bacterioplankton communities may vary between different RAS. Further studies are needed to investigate the effect of reared fish species and aquaculture practices for identification of the key drivers of RAS bacterioplankton communities.
Now that you've reached the end of the article ...
… please consider supporting GSA’s mission to advance responsible seafood practices through education, advocacy and third-party assurances. The Advocate aims to document the evolution of responsible seafood practices and share the expansive knowledge of our vast network of contributors.
By becoming a Global Seafood Alliance member, you’re ensuring that all of the pre-competitive work we do through member benefits, resources and events can continue. Individual membership costs just $50 a year.
Not a GSA member? Join us.
Authors
-
Letícia N. Duarte, MSc.
Department of Biology & CESAM
University of Aveiro, Aveiro, Portugal -
Francisco J.R.C. Coelho, Ph.D.
Department of Biology & CESAM
University of Aveiro, Aveiro, Portugal -
Vanessa Oliveira, Ph.D.
Department of Biology & CESAM
University of Aveiro, Aveiro, Portugal -
Daniel F.R. Cleary, Ph.D.
Department of Biology & CESAM
University of Aveiro, Aveiro, Portugal -
Patrícia Martins, Ph.D.
Department of Biology & CESAM
University of Aveiro, Aveiro, Portugal -
Newton C.M. Gomes, Ph.D.
Corresponding author
Department of Biology & CESAM
University of Aveiro, Aveiro, Portugal[116,112,46,97,117,64,109,99,110,115,101,109,111,103]
Tagged With
Related Posts

Health & Welfare
Calling for a National Broodstock Improvement Network
A National Broodstock Improvement Network for aquaculture could help to break the link between inbreeding and hatchery size.

Responsibility
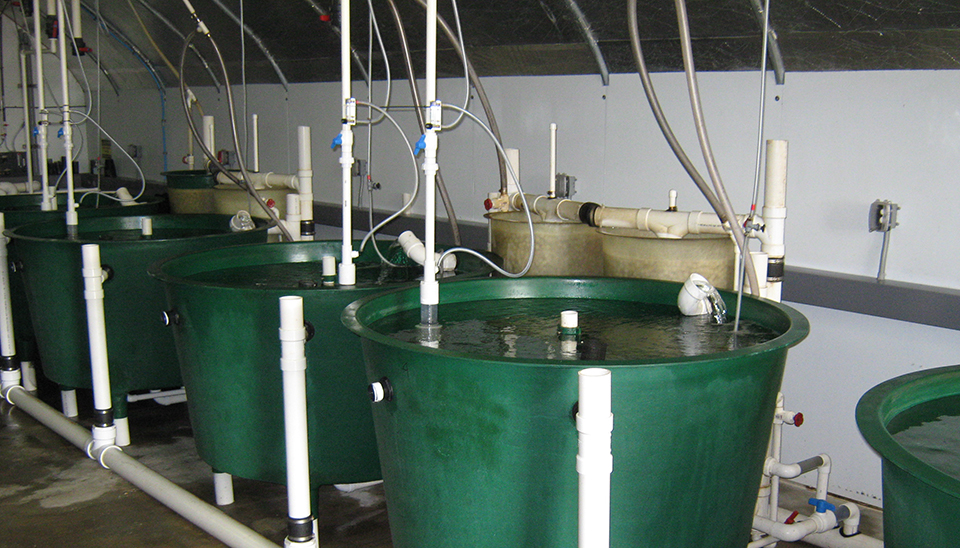
Evaluating low-energy recirculating system for inland production of marine finfish juveniles
U.S. institutions are collaborating to evaluate low-energy recirculating aquaculture system designs to produce marine finfish in a low-salinity environment.

Health & Welfare
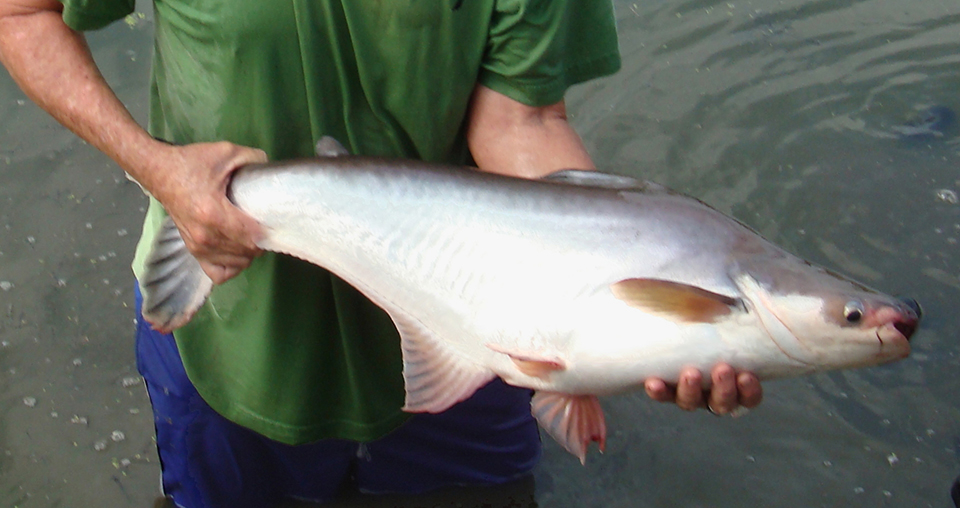
Advances in fish hatchery management
Advances in fish hatchery management – particularly in the areas of brood management and induced spawning – have helped establish aquaculture for multiple species.

Intelligence
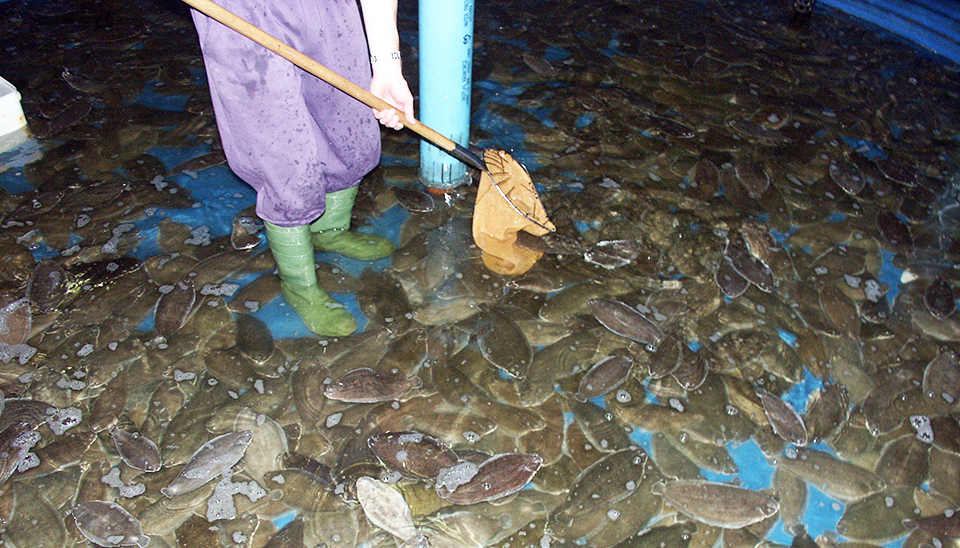
Sole show culture potential in Europe
Early efforts at sole culture in Europe were hampered by several technological and disease problems. Improvements in recirculation technology and advances in feed technology have renewed interest in the species.



