Mini-array diagnostic method offers efficiency, sensitivity
Diagnostic tools that are highly sensitive and specific are needed to manage and prevent shrimp diseases, particularly viral diseases, that have caused losses estimated at billions of dollars. After the development of genomic probe diagnostic systems in the 1990s, followed by polymerase chain reaction- (PCR) based systems, a new mini-array method now allows one-step detection of multiple infections that are common in crustaceans.
The first mini-array colorimetric kit, developed in a collaboration between Skuld Tech and ADIMAQUA of France, provides simultaneous diagnostics for two different viruses: White Spot Syndrome Virus and Infectious Hypodermal and Hematopoietic Necrosis Virus. Both viruses are important because of their wide range and significant impact on shrimp production.
Mini-array method
The mini-array method is based on detection that combines target nucleic acid amplification by PCR and specific hybridization using enzyme-linked antibodies that produce a dark-blue precipitate on a nylon membrane.
This method has numerous advantages over other molecular diagnostic tools, including higher sensitivity, high specificity, greater efficiency and reliability, ease of use, and straightforward interpretation. It eliminates the need for tedious electrophoresis and the use of ethidium bromide, a carcinogenic dye.
Multiple detection possible

Multiple detection is one of the major advantages of mini-arrays. Multiple sets of specific primers are used, and the different amplicons issued from the PCR hybridize with the homologous probes plotted in pre-defined positions on the nylon membrane. Interpretation of results viewed on the membrane can easily be done by the naked eye (Fig. 1).
An internal control is added to each reaction. It is a PCR and hybridization- positive control DNA fragment used to prevent false negatives due to experimental errors. This test is considered positive if the corresponding spots dotted on the membrane show the dark-blue positive staining.
High sensitivity
The mini-array method significantly increases the power detection limit of pathogens and is a highly sensitive diagnostic method that contributes to virus prophylaxis. Its sensitivity is 100 times higher than the regular detection in electrophoresis of PCR-amplified products, and 10,000 times higher than the dot-blot method.
Mini-array is an ideal method for the detection of low-level or early infections. Such early warnings can help operators limit the risks of horizontal transmission by systematically eliminating virus carriers, and the risks of vertical transmission by selecting healthy broodstock individuals.
Conclusion
The mini-array test is fast; between one and 24 reactions can be done in parallel in only six hours. Interpretation of results is easy, as the presence or absence of blue dots can be detected by the naked eye. Sample processing is also easy, involving simple heating in boiling water.
The membranes provided with the mini-array kits are ready to use, dotted with the probes, and pre-hybridized. Only a few items of equipment are required. Except for the thermocycler, only basic equipment such as a water bath, small bench centrifuge and shaker are necessary. Thus, the mini-array method can be easily implemented at shrimp farms or hatcheries that already use PCR as a diagnostic tool.
(Editor’s Note: This article was originally published in the December 2000 print edition of the Global Aquaculture Advocate.)
Now that you've reached the end of the article ...
… please consider supporting GSA’s mission to advance responsible seafood practices through education, advocacy and third-party assurances. The Advocate aims to document the evolution of responsible seafood practices and share the expansive knowledge of our vast network of contributors.
By becoming a Global Seafood Alliance member, you’re ensuring that all of the pre-competitive work we do through member benefits, resources and events can continue. Individual membership costs just $50 a year.
Not a GSA member? Join us.
Author
-
Jean Robert Bonami, Ph.D.
ADIMAQUA
Montpellier, France[114,102,46,50,112,116,110,111,109,45,118,105,110,117,46,116,105,114,99,64,105,109,97,110,111,98]
Tagged With
Related Posts

Health & Welfare
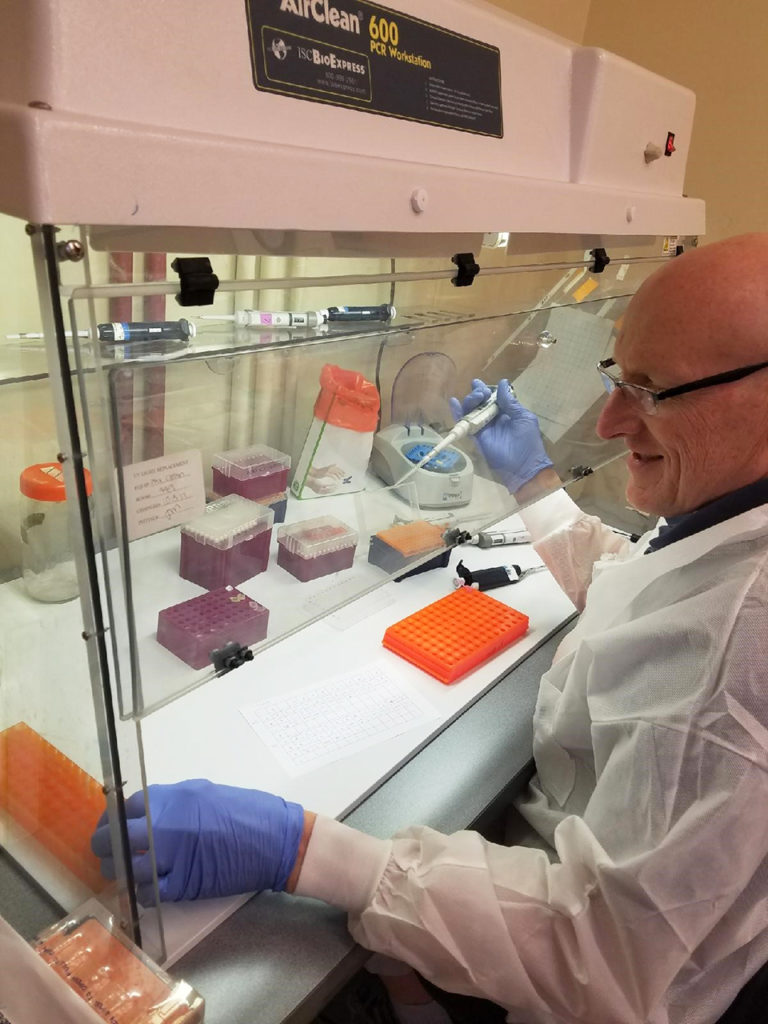
A comprehensive look at the Proficiency Test for farmed shrimp
The University of Arizona Aquaculture Pathology Laboratory has carried out the Proficiency Test (PT) since 2005, with 300-plus diagnostic laboratories participating while improving their capabilities in the diagnosis of several shrimp pathogens.

Health & Welfare
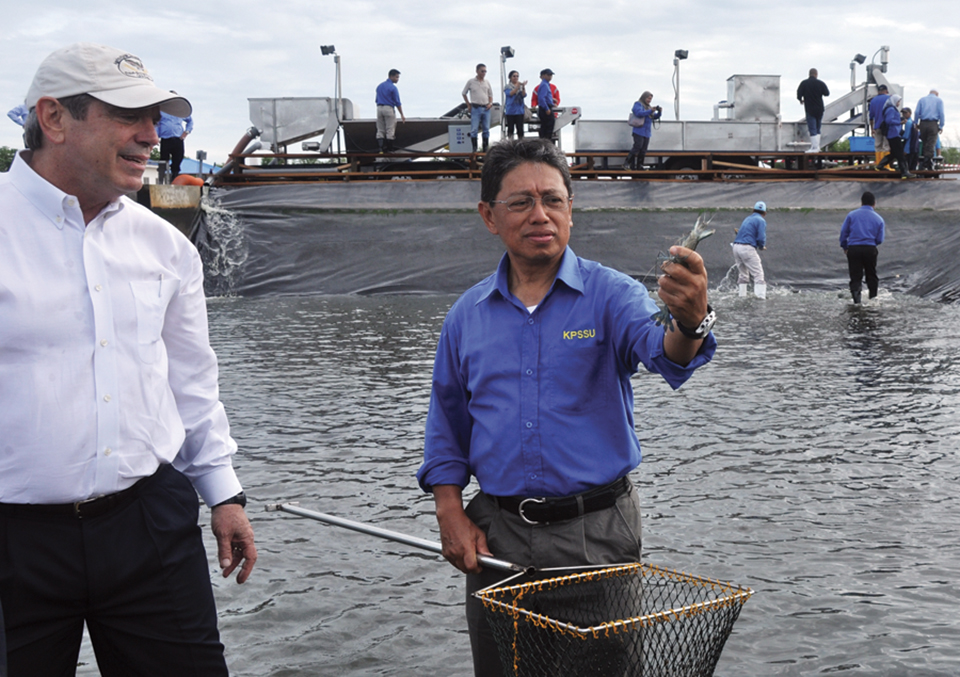
Brunei project develops technology for large black tiger shrimp production, part 1
A five-year project was undertaken in Brunei Darussalam to develop advanced technology for the production of large black tiger shrimp. A combination of technologies has enabled efficient production of large-sized black tiger shrimp, which could lead to a resurgence of this species in Asia.

Health & Welfare
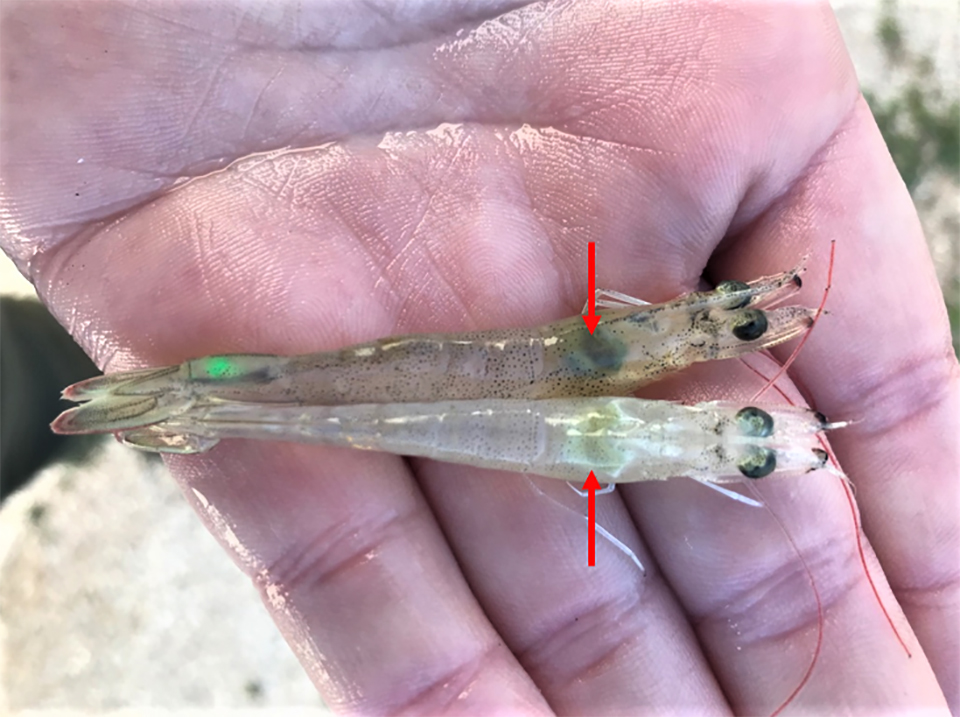
AHPND is a chronic disease in Pacific white shrimp from Latin America
There is a new phase of infection for Acute Hepatopancreatic Necrosis Disease on Latin American shrimp farms, a contrast to observations in Southeast Asia.

Health & Welfare
The unmet promise of pondside PCR
A new generation of technology, portable PCR, offers potential for affordable, immediate, pondside diagnosis in easy-to-use handheld kits. But will it live up to the hype?



