Shrimp disease originated in Ecuador
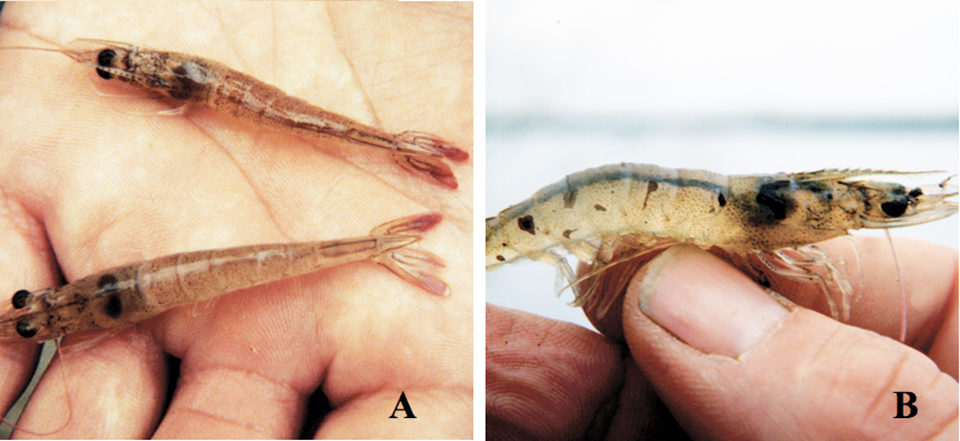
Taura syndrome virus (TSV) is a major pathogen of Litopenaeus vannamei that causes mortalities of 40 to 95 percent in farmed shrimp populations and results in substantial economic losses. The virus was first reported from Ecuador in 1992, but is now widely distributed throughout the Americas and Southeast Asia.
Study setup
To determine the evolutionary history of this virus, the authors amplified and sequenced the viral capsid protein 2 gene (CP2) among 83 TSV isolates collected from 1992 to 2008 in 16 countries. They performed a phylogenetic analysis using the Bayesian Evolutionary Analysis Sampling Trees (BEAST) program with these sequences.
The results revealed four major TSV lineages – designated Mexico, Southeast Asia, Belize and Venezuela – as well as the position of the root in the phylogenetic tree (Fig. 1). The most basal lineage included a cluster of strains isolated in Colombia, Ecuador and the United States. This finding corresponded with the first recognition and description of Taura syndrome in Ecuador in 1991.
Mexico lineage
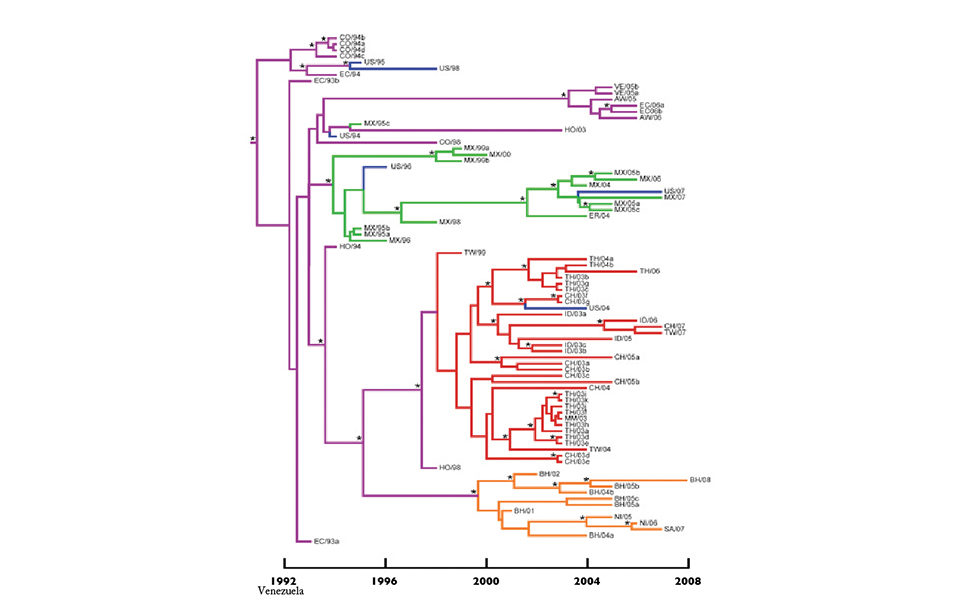
The first major lineage consisted of isolates collected in Mexico. Thirteen of the 14 Mexican isolates analyzed were within this group. Therefore, the majority of Mexican TSV outbreaks were likely the result of a single introduction to the country. This lineage also clustered with TSV found in Eritrea and in Texas, USA, in 1996 and Hawaii, USA, in 2007. It is suspected these later outbreaks stemmed from imports from Mexico.
Southeast Asia lineage
The second major TSV lineage was from Southeast Asia and possibly originated in Honduras. The first case was reported in vannamei in Taiwan in 1998, and a Taiwan isolate (TW/99) clustered with a Honduran isolate (HO/98). At that time, this shrimp species was not found in Asia but is indigenous to the Americas, strengthening the case that TSV was probably exported to Taiwan from Honduras.
Within this lineage, TSV isolated in China, Taiwan and Thailand was dispersed among each other, suggesting multiple transmission events among these three countries. TSV strains from Indonesia formed a particularly strong group and may have been the result of a single introduction into that country, because all the Indonesian TSV descended from a single common ancestor. Later, Indonesian TSV was apparently transmitted back to Taiwan and China.
Belize lineage
The third major TSV lineage was from Belize. This strain is known to be highly virulent, and isolates from Belize formed a distinct group together with isolates from Nicaragua (NI/05 and NI/06), which is nearby, and Saudi Arabia (SA/07). The strain was first found in 2001 and was still present in Belize in 2008, indicating it had not been eradicated from local farms.
The reports of isolates from Nicaragua and Saudi Arabia indicated that shrimp from producers in Belize was shipped among these countries. In these cases, similar to the situations in Mexico and Southeast Asia, shrimp farmers were moving stocks between facilities without routine disease monitoring.
Venezuela lineage
Finally, a phylogenetically distinct TSV group comprised of Venezuela and Aruba isolates was inferred. The Venezuela/Aruba TSV lineage was separated from the other TSV isolates by a long internal branch, suggesting a distinct epidemic.
The Venezuela and Aruba isolates were from two farms owned by the same company, which frequently shipped broodstock and postlarvae between the two facilities.
Taura syndrome in U.S.
In the U.S., where disease monitoring has been more rigorous, TSV outbreaks have been sporadic. Outbreaks occurred in 1994, 1995, 1996, 1998, 2004 and 2007 in Texas or Hawaii. Of these six isolates, only two were closely related: Hawaii in 1995 (US/95) and Texas in 1998 (US/98). These two isolates were possibly the result of the same transmission event into the U.S., but were separated by a three-year interval, indicating the 1998 Texas outbreak did not stem from the earlier outbreak in Hawaii.
The cases probably both originated from Ecuador, since both clustered with a 1994 isolate from that country. Another two isolates from Texas (US/96) and Hawaii (US/07) fell within the Mexican lineage. A Texas isolate (US/04) was grouped within the Southeast Asia lineage.
Since the first TSV isolated in Hawaii (US/94) fell within an unresolved portion of the phylogenetic tree, the precise geographical location was unknown. The phylogenetic distance separating the majority of TSV isolates from the U.S. indicates that the monitoring and containment protocols employed by the U.S. shrimp aquaculture industry are highly effective at detecting outbreaks and preventing their further spread.
Mutation rate
The phylogenetic analysis also provided insight into the rate of TSV evolution. The TSV substitution rate of 2.37 x 10-3 substitutions/site/year is similar to rapidly evolving RNA viruses such as swine vesicular disease, human enterovirus 71 and human immunodeficiency virus type 1.
The high rate of substitution may explain the increasing diversity of TSV associated with its spread among geographic regions. This, in turn, may result in the need for perpetual development of new TSV reverse transcription polymerase chain reaction primers and continued searches to find new populations of TSV-resistant P. vannamei.
(Editor’s Note: This article was originally published in the September/October 2009 print edition of the Global Aquaculture Advocate.)
Now that you've reached the end of the article ...
… please consider supporting GSA’s mission to advance responsible seafood practices through education, advocacy and third-party assurances. The Advocate aims to document the evolution of responsible seafood practices and share the expansive knowledge of our vast network of contributors.
By becoming a Global Seafood Alliance member, you’re ensuring that all of the pre-competitive work we do through member benefits, resources and events can continue. Individual membership costs just $50 a year.
Not a GSA member? Join us.
Authors
-
Kathy F.J. Tang, Ph.D.
Department of Veterinary Science and Microbiology
University of Arizona
Tucson, Arizona 85721 USA[117,100,101,46,97,110,111,122,105,114,97,46,117,64,117,121,106,103,110,101,102]
-
Joel O. Wertheim
Department of Veterinary Science and Microbiology
University of Arizona
Tucson, Arizona 85721 USA -
Solangel A. Navarro
Department of Veterinary Science and Microbiology
University of Arizona
Tucson, Arizona 85721 USA -

Donald V. Lightner, Ph.D.
Department of Veterinary Science and Microbiology
University of Arizona
Tucson, Arizona 85721 USA
Tagged With
Related Posts
Health & Welfare
New WSSV, TSV genotypes identified in Saudi Arabia
To determine origins of White Spot Syndrome Virus and Taura Syndrome Virus in Saudi Arabia, authors performed genotyping studies and found a new genotype in each of the isolates collected.

Health & Welfare
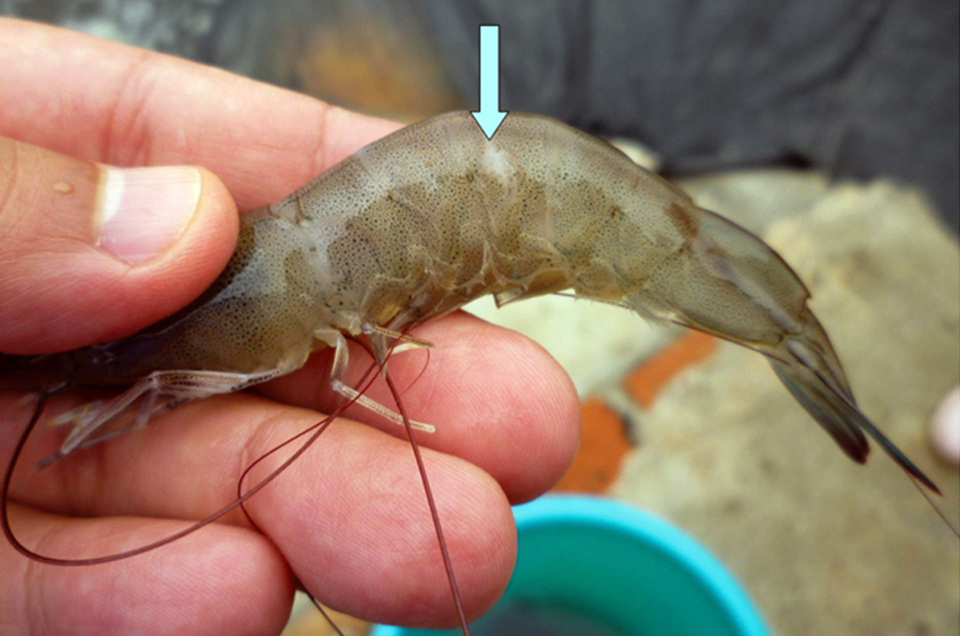
New procedure controls TSV in white shrimp
The authors have established a procedure to reduce the impacts of Taura Syndrome Virus in the culture of Pacific white shrimp. The procedure focuses on avoiding the molting process of shrimp by limiting culture conditions.

Health & Welfare
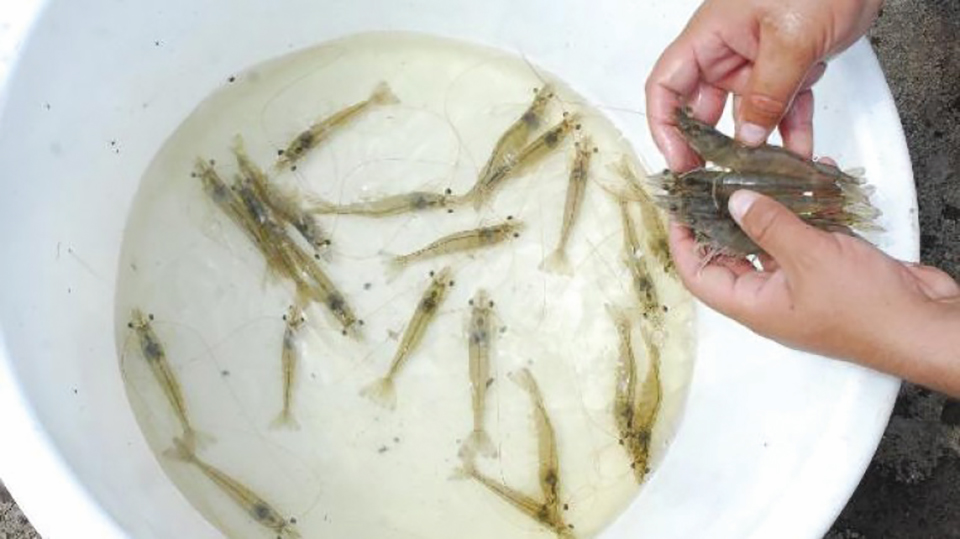
A study of Zoea-2 Syndrome in hatcheries in India, part 1
Indian shrimp hatcheries have experienced larval mortality in the zoea-2 stage, with molt deterioration and resulting in heavy mortality. Authors investigated the problem holistically.

Health & Welfare
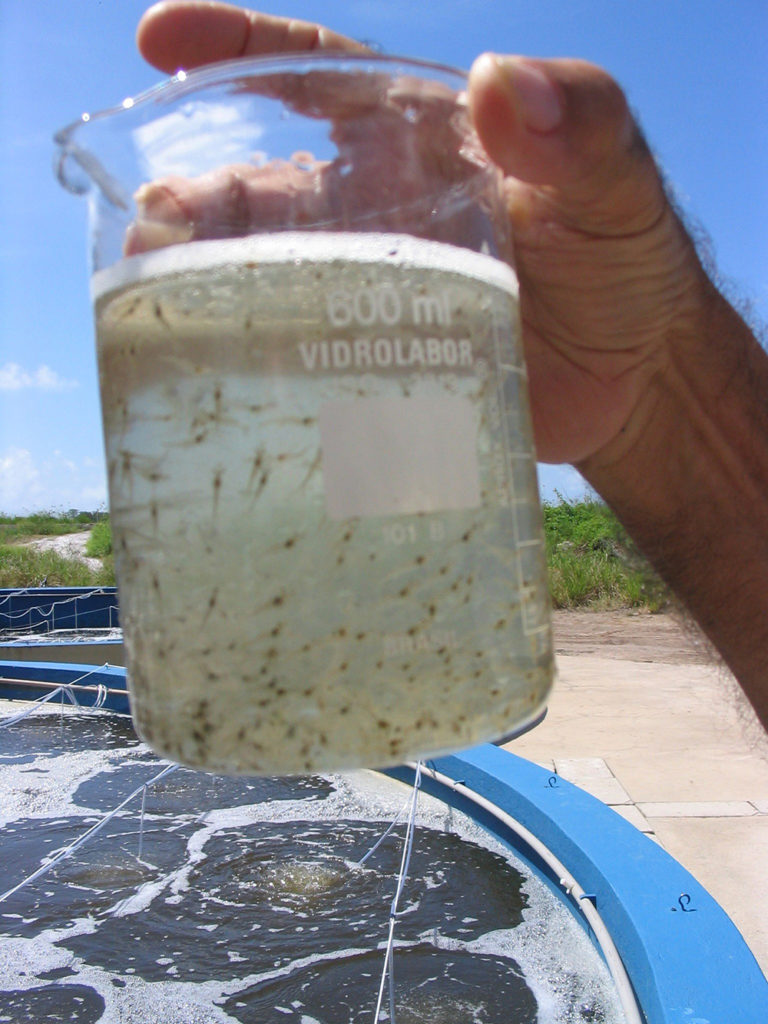
A study of Zoea-2 Syndrome in hatcheries in India, part 2
Indian shrimp hatcheries have experienced larval mortality in the zoea-2 stage, with molt deterioration and resulting in heavy mortality. Authors considered biotic and abiotic factors. Part 2 describes results of their study.


