Emerging solution for resistance against diseases like enteric septicemia
Channel catfish (Ictalurus punctatus) production in the United States has developed over the past 40 years to establish the species as the primary foodfish of the U.S. aquaculture industry. The U.S. Department of Agriculture’s Agricultural Research Service (USDA-ARS) has developed a selective-breeding program for these important fish and recently released a genetically improved NWAC103 strain to the industry in cooperation with Mississippi State University. Research on subsequent generations of this strain will result in further improvement of economically important traits, including improved disease resistance, in the fish.

Enteric septicemia of catfish (ESC) is the most prevalent and costly disease affecting channel catfish commercial production. Caused by the bacterium Edwardsiella ictaluri, ESC occurs most commonly in the fall and spring, when water temperatures are 18 to 27 degrees-C.
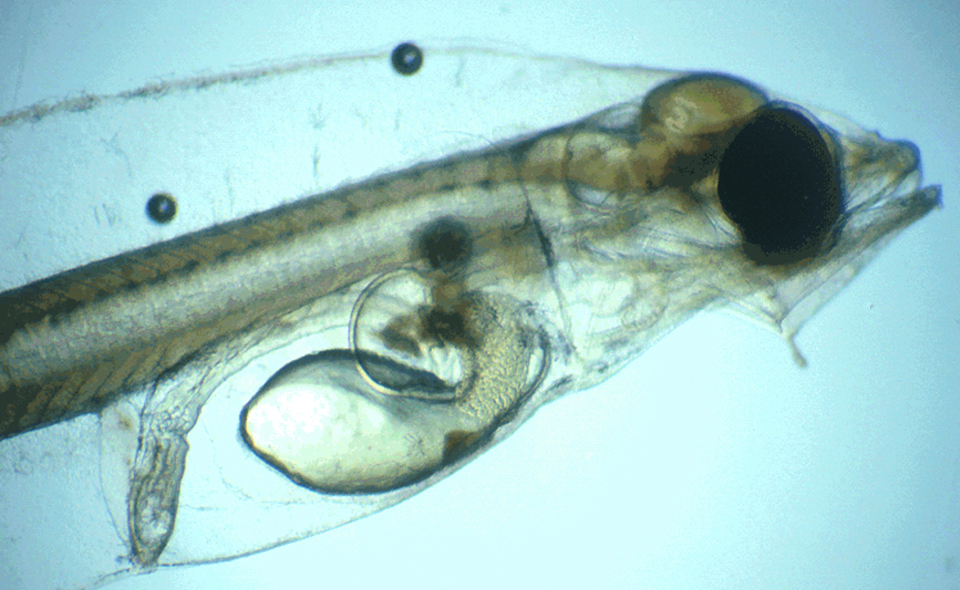
Pathogen uptake occurs primarily through the gut, nares and gills and appears to be facilitated by active feeding during periods of disease outbreak. Progression of ESC is usually rapid and most significantly affects fingerlings. Clinical disease signs that manifest within a few days post-infection (Fig. 1) are usually followed by death within two weeks. A better understanding of the early stages of ESC infection and clearance of E. ictaluri in exposed fish could improve selective breeding for disease resistance.
Pathogen detection
Sensitive detection of ESC is important to the successful development of a selective-breeding program for ESC resistance. Until recently, however, accurate quantitative assessment of pathogen levels has been problematic when infection levels are low.
Therefore, a genetic assay using quantitative polymerase chain reaction was developed for early detection of E. ictaluri DNA in channel catfish blood. The assay provides high accuracy and specificity and is capable of detecting as few as 2.5 cells of E. ictaluri in 100 μl of infected blood in less than three hours from the time of sample collection.
Use in genetic improvement
When applied to selective breeding for disease resistance, the genetic assay provides the tools necessary to track the early stages of infection and the sensitivity necessary to detect differences among fish populations per families that have different levels of innate disease resistance. The assay can also be used to further study the kinetics of ESC infection and monitor clearance rates during infections.
Disease resistance
NWAC103 channel catfish families show a consistent response to ESC infection relative to susceptibility. But some families are highly resistant to ESC and others are highly susceptible (Fig. 2). Within each family there is a large degree of variability in response to the infection.
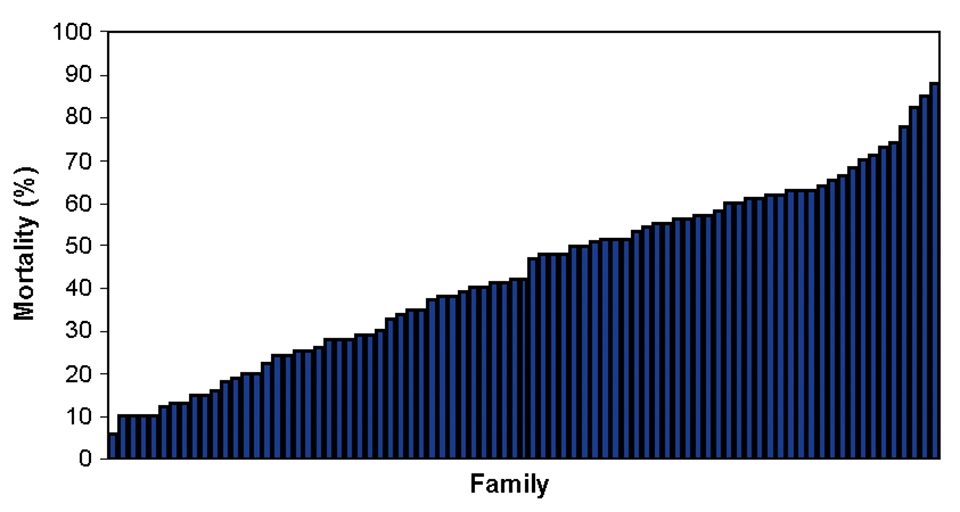
This complicates selective breeding for disease resistance. Acquiring specific information on disease progression at early stages of infection could help by having a trait in addition to mortality to utilize as a selection criterion.
Bacterial levels and clearance
A spring 2002 study by USDA-ARS scientists at the Thad Cochran National Warmwater Aquaculture Center in Stoneville, Mississippi, USA addressed how bacterial levels and disease clearance vary in experimental channel catfish families. Two families that showed a consistent response to ESC in prior experiments – one susceptible and the other resistant – were challenged with E. ictaluri. Initial uptake of the pathogen, changes in bacterial load over time, and clearance of the pathogen from the animals’ blood were measured with the genetic detection assay.
Initial uptake was assessed two hours after fish were exposed to E. ictaluri. No difference was evident based on bacterial DNA levels in the blood samples. This indicated that any differences in susceptibility were not related to uptake of the bacteria.
One to five days after exposure, fish from the susceptible family carried higher bacterial levels in their blood than fish from the resistant family (Fig. 3). However, when the rate of decrease in bacterial concentrations was measured, there was no difference between the groups (Fig. 4).
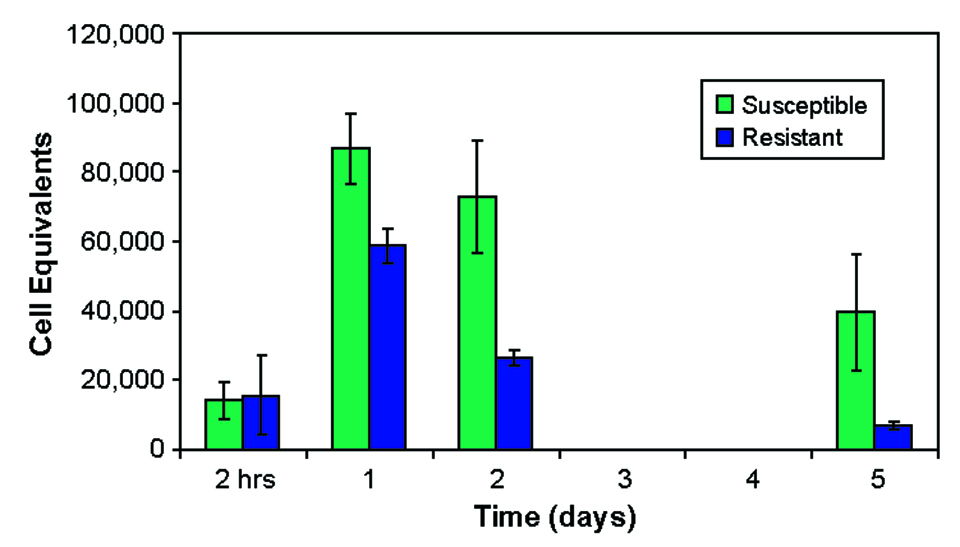
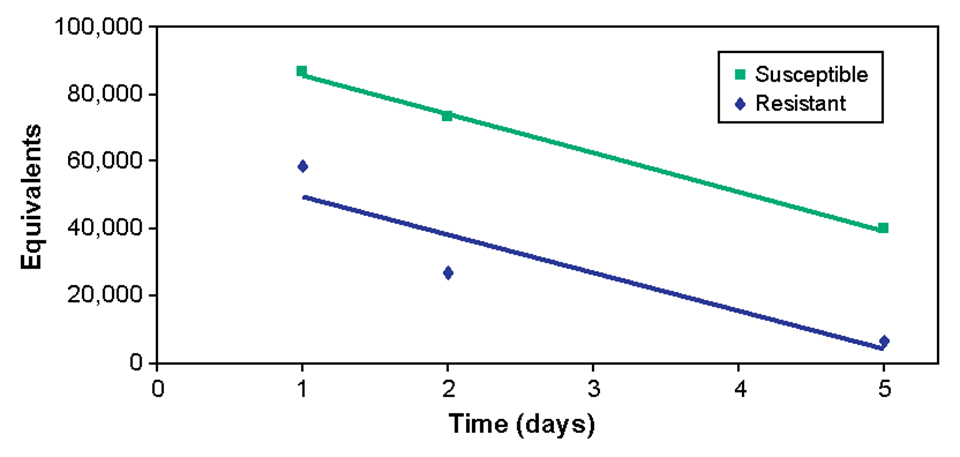
This suggested that survival after ESC infection is not determined by clearance rates from blood and that the immune system can suppress ESC infection in resistant fish. Continued low-level infections in resistant fish may be a source of subsequent infections of ESC.
Conclusion
The genetic detection assay described here will improve selective-breeding programs by providing an assessment of early stages of infection and an understanding of the mechanisms responsible for disease resistance. In this case, comparison of resistant and susceptible fish showed that initial uptake did not affect survival, that susceptible fish had higher levels of bacteria 24 hours after exposure compared to resistant fish, and that clearance of E. ictaluri was equivalent for both susceptible and resistant fish.
(Editor’s Note: This article was originally published in the February 2003 print edition of the Global Aquaculture Advocate.)
Now that you've reached the end of the article ...
… please consider supporting GSA’s mission to advance responsible seafood practices through education, advocacy and third-party assurances. The Advocate aims to document the evolution of responsible seafood practices and share the expansive knowledge of our vast network of contributors.
By becoming a Global Seafood Alliance member, you’re ensuring that all of the pre-competitive work we do through member benefits, resources and events can continue. Individual membership costs just $50 a year.
Not a GSA member? Join us.
Authors
-
Lelania Bilodeau, Ph.D.
USDA-ARS Catfish Genetics Research Unit
Thad Cochran National Warmwater Aquaculture Center
141 Experiment Station Road
Stoneville, Mississippi 38776 USA[118,111,103,46,97,100,115,117,46,115,114,97,64,117,97,101,100,111,108,105,98,97]
-
William R. Wolters, Ph.D.
USDA-ARS Catfish Genetics Research Unit
Thad Cochran National Warmwater Aquaculture Center
141 Experiment Station Road
Stoneville, Mississippi 38776 USA -
David J. Wise, Ph.D.
Delta Research Extension Center
Mississippi State University
Thad Cochran National Warmwater Aquaculture Center
Stoneville, Mississippi, USA
Tagged With
Related Posts

Health & Welfare
Black tiger domestication, selective breeding advance in Australia
Using clear-water tank systems, CSIRO and a collaborating farm have advanced the domestication of black tiger stocks in Australia.

Health & Welfare
10 paths to low productivity and profitability with tilapia in sub-Saharan Africa
Tilapia culture in sub-Saharan Africa suffers from low productivity and profitability. A comprehensive management approach is needed to address the root causes.

Intelligence
Adding value to tilapia to tap into U.S. market
New markets for tilapia and expansion of existing ones can be created by planning and implementing properly designed geographic strategies to meet discriminating consumer preferences. Low labor costs in most producing countries promotes value-adding by the production of fresh fillets.
Health & Welfare
Scope for further optimization of selective breeding in aquaculture species
Article briefly describes the structure of family-based selective breeding and identifies areas where developments in genetics and related fields may increase efficiency.


