Research comes at a high cost
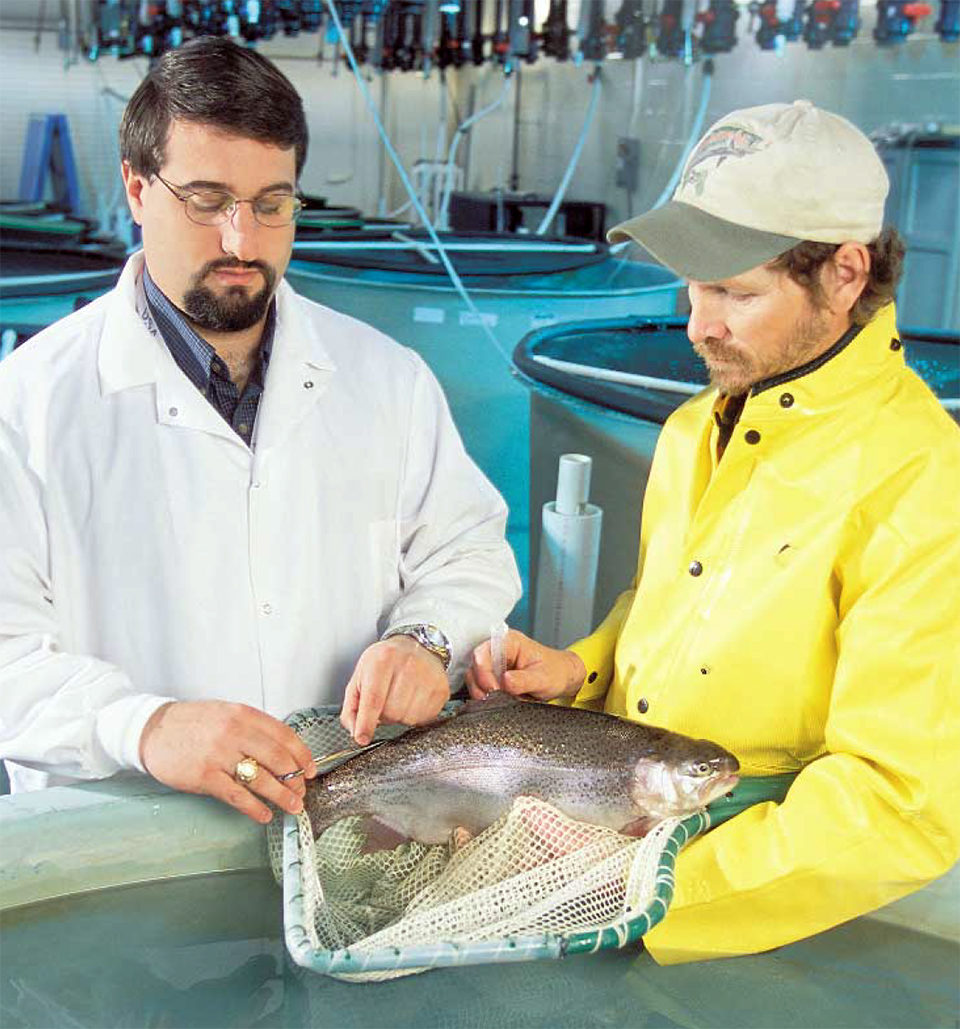

Recent advances in aquaculture research include the integration of genomics – the study of genes and their functions – into projects pursuing increased aquaculture production efficiency. The genetic material or genome of every species contains the blueprint that guides its biological processes. Genome research can, therefore, provide understanding of the mechanisms that drive these processes and provide knowledge to manipulate them for desired outcomes.
Salmon genomics
The primary goal of salmonid genomics as it relates to aquaculture is to identify the genes that affect production traits and integrate that information into selective-breeding programs aimed at genetic improvement. Desired outcomes include the development of strains which are disease-resistant, stress-tolerant, fast-growing, and reproductively manageable.
To this end, genome tools and technologies must be integrated with other scientific disciplines involved in the characterization of traits, such as quantitative genetics and the study of physiology, fish health, and production systems.
https://www.aquaculturealliance.org/advocate/aquagen-ceo-genomics-are-transforming-aquaculture/
Genetic doubling
One of the special difficulties for genome research in salmonids involves their evolutionarily recent genome duplication about 100 million years ago. In this event, the amount of genetic material doubled to produce an organism genetically different from its ancestors.
Initially this resulted in two copies of each chromosome and therefore each gene. Over time, chromosomes and gene sequences have diverged to become different from the ancestral versions and one another. Although the gene sequences continue to diverge throughout time, they typically perform similar functions, often in the same tissues at the same time. This complicates genetic studies, as researchers might assume they are studying the effects of a single gene but in reality may be dealing with two or more.
Ordinarily, engaging in genome research requires the development of species-specific resources. However, due to the recent evolutionary divergence of the salmonids, many of the tools developed for one species are useful for others. For instance, many of the microsatellite genetic markers developed in Atlantic salmon also work for rainbow trout and Artic char, and vice versa.
This genome similarity is often exploited, as genome tools and reagents are very expensive and time-consuming to develop. For this reason, genome projects are typically undertaken by whole academic communities consisting of multiple university, government, and industry laboratories, and include international collaborations. Limited amounts of comparative genome information can be obtained from well-studied species such as zebrafish, pufferfish, and to a lesser extent, humans and mice.
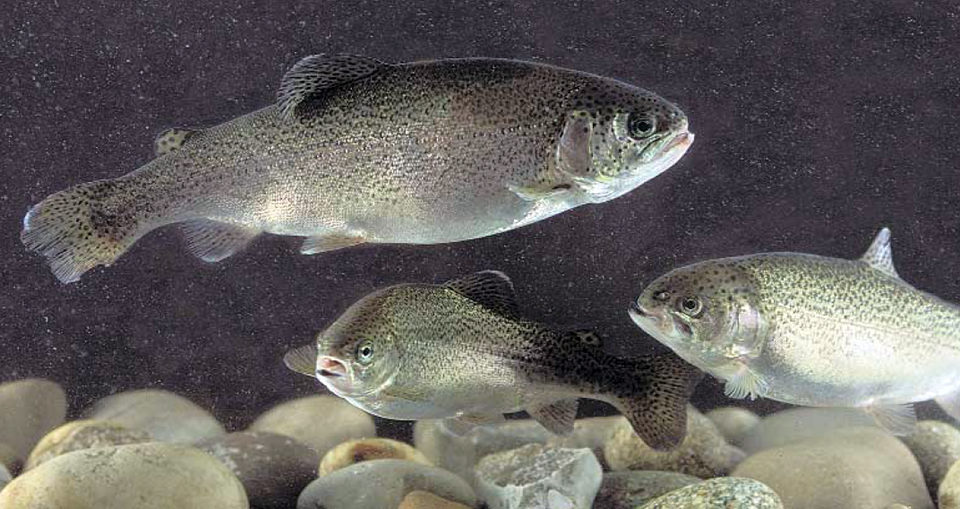

Functional genomics, proteomics
The central dogma of molecular biology, first stated by Francis Crick in 1956, describes the flow of information in a cell. Briefly, DNA molecules are transcribed into RNA molecules, which are then translated into protein molecules, which perform specific biological functions. Genome research must, therefore, be heavily integrated with functional genomics and proteomics.
Functional genomics focuses on RNA transcription, more specifically in what tissues and at what stage in life specific RNA sequences are transcribed. Proteomics centers on the presence of proteins and their functions.
Currently, resources are being developed to study the genome, the transcriptome, and the proteome in a high-throughput manner that allows the observation of many genes in single experiments. Information on all three of these molecules can be correlated with performance data to identify genes that affect production traits.
Genome tools
Tools of the genome trade can be categorized as sequence information or chromosome maps. Sequence information can be derived from DNA, RNA, and protein sequences. All of these sequences are available in GenBank, a public database maintained by the U.S. National Center for Biotechnology Information, part of the National Institutes of Health.
Over the last two years, there has been a dramatic increase in the amount of sequence information for salmonids. The majority of this information – transcriptome sequences for rainbow trout and Atlantic salmon – comes from projects aimed at gene discovery.
Physical maps
Chromosome maps fall into one of two subcategories: physical maps or genetic maps. Most physical maps contain information about the physical structures of chromosomes in terms of chromosome number, shape, locations of genes, and comparisons to chromosomes from other species. Determination of the full genome sequences for these species would provide the ultimate physical map and sequence information, and greatly enhance our ability to accomplish research in these species.
For example, rather than many fish health laboratories spending years to identify and sequence immune-related genes in the laboratory, the whole genome sequence would allow for gene identification in silico by computation. This would greatly accelerate discoveries in this area. International consortiums are currently working to obtain funding to sequence the entire genomes of rainbow trout and Atlantic salmon.
Genetic maps
Genetic maps contain information about how genetic markers are inherited. Genetic markers such as micro-satellites are most often random sequences of DNA, but sometimes are associated with a gene. Correlating marker inheritance with traits of importance can give clues to the genes that affect that trait.
Genetic markers have been correlated with salinity tolerance, temperature tolerance, albinism, spawning time, embryonic development rate, and viral disease resistance in rainbow trout.
To move from genetic markers to identification of the genes affecting the trait requires a large research effort utilizing physical maps, genetic maps, and DNA, RNA, and protein sequence information. This information can then be integrated into selective-breeding programs through marker-assisted selection. Genetic maps have been created or are currently under construction for rainbow trout, Atlantic salmon, Arctic char, and Chinook salmon.
Conclusion
Genome research includes sets of tools and information for use in studying the characteristics of organisms. Salmonid genome research has recently come out of its infancy with the development of species-specific resources.
As researchers move forward in the identification of genes that affect aquaculture production traits, it will be extremely important to use commercially relevant germplasm in research experiments to facilitate the incorporation of molecular data into commercial broodstock programs. Although the applications of genome research have great potential, the high cost of this research and the two- to four-year generation interval of these species are impediments to an immediate impact on the aquaculture industry.
(Editor’s Note: This article was originally published in the April 2004 print edition of the Global Aquaculture Advocate.)
Now that you've reached the end of the article ...
… please consider supporting GSA’s mission to advance responsible seafood practices through education, advocacy and third-party assurances. The Advocate aims to document the evolution of responsible seafood practices and share the expansive knowledge of our vast network of contributors.
By becoming a Global Seafood Alliance member, you’re ensuring that all of the pre-competitive work we do through member benefits, resources and events can continue. Individual membership costs just $50 a year.
Not a GSA member? Join us.
Author
-
Caird E. Rexroad III, Ph.D.
USDA/ARS National Center for Cool and Cold Water Aquaculture
11861 Leetown Road
Kearneysville, West Virginia
25430 USA[118,111,103,46,97,100,115,117,46,115,114,97,46,97,119,99,99,99,110,64,100,97,111,114,120,101,114,99]
Tagged With
Related Posts

Innovation & Investment
AquaGen CEO: Genomics are transforming aquaculture
The CEO of AquaGen knew that the Norwegian research group’s work in genomics was key to the salmon industry’s future. And that was before she even worked there.

Health & Welfare
Atlantic cod genomics and broodstock development project
The Atlantic Cod Genomics and Broodstock Development Project has expanded the gene-related resources for the species in Canada.

Health & Welfare
Advances in tilapia nutrition, part 2
Nutrition has an important role on growth, performance and flesh quality of tilapia. Part two of this two-part series looks into mineral supplementation and feeding strategy.
Health & Welfare
Animal health giants have sea lice in their crosshairs
Alltech and Benchmark have been working on the next generation of sea lice solutions and believe they have new products that can help salmon farmers win.


