Genetic selection for viral resistance
Classical genetics is based on heritable variations in individuals that allow one to deduce the existence of underlying elements called genes. The existence of a gene is thus known as a result of mathematical deduction. In contrast, modern (molecular) genetics is based on examination of the hereditary molecules themselves – the organism’s DNA.
Molecular genetics has been used extensively in terrestrial agriculture, especially to improve crops and animals of high economic value. The application of modern genetic technology to aquaculture, however, is in its infancy. The size of the aquaculture industry, the demands placed on it by an expanding world population, and the threat of uncontrolled disease demand implementation of improved methods of husbandry that include the application of modern genetic technology.
Molecular markers
Molecular genetics for plant and animal breeding is based on the use of molecular markers that can be used as signposts to identify the presence or absence of genes responsible for traits of interest. Such markers are elements in the DNA that are co-inherited with genes. Identification of the markers associated with a gene substitutes for direct identification of an otherwise unidentifiable gene responsible for a trait.
Several classes of markers have been developed over the past few decades. Each has advantages and disadvantages. Microsatellites, a type of molecular marker that consists of repeating sequences of nucleic acids, are commonly chosen for many applications.
Isolating microsatellites
Isolation of suitable markers requires considerable effort. Microsatellites are often isolated through an enrichment procedure that selectively removes microsatellite- containing DNA fragments from a total mixture of fragments and incorporates them into bacterial cells as recombinant DNA. This is a technically complex process, but once experience has been gained, enriched libraries can be produced within a matter of weeks.
Only about 10 percent of the DNA sequences obtained from both L. vannamei and P. monodon result in usable microsatellites, so many more need to be isolated than ultimately are utilized. Enriched libraries have been produced from L. vannamei that contain many thousands of microsatellites, providing a source of potential markers for use in such diverse applications as stock identification, parent/offspring and relatedness determination, diversity monitoring, and marker-assisted selection for traits of economic importance to the industry.
Allelic diversity
Most genes exist in a variety of forms, and each variant is referred to as an allele. The variability of many traits is due to variability in the alleles that control the trait. Genetic theory predicts that inbreeding will lead to a loss of alleles, which in turn results in a decrease in genetic fitness of a stock. A stock’s ability to respond to stress and perhaps its ability to withstand various diseases is correspondingly diminished.
For that reason, it is desirable to maintain as much allelic diversity as possible, especially if the stock is being subjected to selection for a particular trait. Microsatellites are ideal for monitoring diversity and providing information necessary to offset loss that will otherwise occur during periods of intense selection. For example, in a study using only 10 microsatellites, a stock of L. vannamei bred at random for 20 generations was shown to have approximately 58 percent less diversity, as measured by the average number of alleles that remained in the stock after 20 generations of random mating.
Genetic diversity
The significance of this reduction in diversity was demonstrated by the results of a Taura Syndrome virus (TSV) challenge conducted by Dr. Don Lightner in Arizona, USA. Farmed animals were challenged with a diet of TSV-positive shrimp flesh. Table 1 shows the differences observed between animals that succumbed to the challenge and those that survived.
Using microsatellites, the survivors were shown to contain an average of about 13 percent more alleles. Looked at another way, the chance of any two animals picked at random being identical with respect to these microsatellites was 120 times less in the survivors than in the non-survivors. This strongly suggests that survival depends to a significant extent on the genetic makeup of an animal, and shows that the animals that survived were, as a group, considerably more diverse than those that died.
WSSV research in Panama
In challenges carried out by Farallon Aquaculture of Panama, approximately 200 million postlarvae (produced from 10,000 wild broodstock) were fed flesh positive for WSSV. Subsequently, moribund postlarvae (designated M1) were separated from presumptive survivors that still behaved normally (designated S1). S1 animals were again fed WSSV-positive shrimp flesh and separated into moribund (M2) and survivor (S2) categories. Twice-challenged S2 survivors represented one of every 80,000 of the original animals. Samples representing M1 and S2 groups were then used for genotyping and marker comparison.
Microsatellites as markers
There was no obvious difference between the moribund and surviving animals with respect to genetic diversity. This is probably because the animals were of wild stock origin and the overall high level of genetic diversity masked genetic differences in the two groups. When the microsatellite composition of each of the surviving animals was compared with that of the moribunds, however, the majority of them were found to be more similar to the S2 group than to the M2 group.
This demonstrates that there was, in fact, a genetic difference between the groups, however subtle. Upon closer examination, six microsatellites were found that showed significant allelic frequency differences between the moribund and survivor groups. These microsatellites are thus candidate markers to use in a broodstock selection program to produce offspring that display enhanced resistance in the presence of WSSV.
Testing for WSSV
The polymerase chain reaction (PCR) was used to detect the presence of WSSV in the challenged animals. WSSV primers developed and used widely in Thailand were applied to known infected and non-infected tissue of P. monodon, which served as positive and negative controls. As expected, these produced positive and negative readings, respectively.
These same PCR primers were then used on M1 L. vannamei and this entire group tested positive. None of the animals from the twice-challenged S2 group, however, tested positive for WSSV. Perhaps their genetic composition reduced their susceptibility to ingested viral material, which is probably also the primary means of infection within a pond environment.
Conclusion
Genetic selection for viral resistance or survivability is an appropriate trait upon which to focus initially because of the devastating effects of viral disease on the industry. There are many other economically important traits in shrimp that have a genetic basis. Once viral disease has been brought under control, microsatellites used in markerassisted selection for any of these traits can lead to general stock improvement. Over the next several years, the shrimp industry should benefit enormously from improvements made possible by this class of genetic markers.
(Editor’s Note: This article was originally published in the December 2000 print edition of the Global Aquaculture Advocate.)
Now that you've reached the end of the article ...
… please consider supporting GSA’s mission to advance responsible seafood practices through education, advocacy and third-party assurances. The Advocate aims to document the evolution of responsible seafood practices and share the expansive knowledge of our vast network of contributors.
By becoming a Global Seafood Alliance member, you’re ensuring that all of the pre-competitive work we do through member benefits, resources and events can continue. Individual membership costs just $50 a year.
Not a GSA member? Join us.
Author
-
Kenneth Jones, Ph.D.
Aquatic Stock Improvement Co.
Hawthorne, California USA[109,111,99,46,115,114,101,107,114,97,109,99,105,116,97,117,113,97,111,99,105,115,97,64,111,99,105,115,97]
Tagged With
Related Posts

Health & Welfare
DNA fingerprinting: System for black tiger shrimp
Molecular markers can allow on-farm selection of high-performance families without the need to use costly alternatives such as separate rearing of families/ groups of families or tagging animals.

Health & Welfare
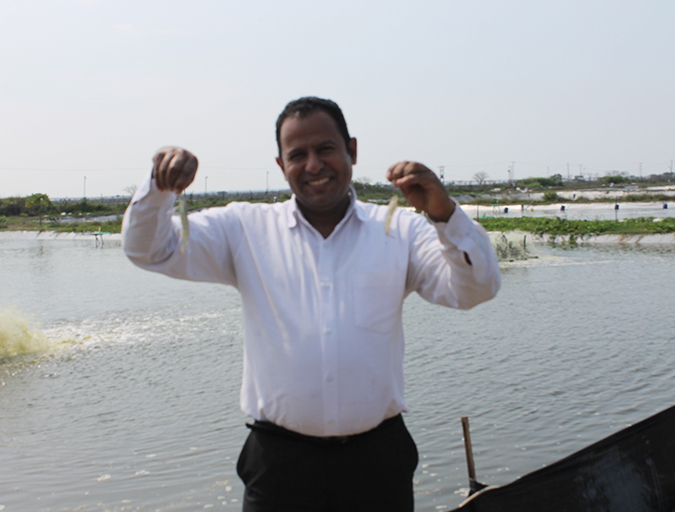
Selection for disease resistance in Indian white shrimp
A new breeding program for genetic improvement of Indian white shrimp (Fenneropenaeus indicus) in Egypt was established at a private shrimp farm at AL Dibah Triangle Zone (DTZ) to develop disease-resistant shrimp for the country’s aquaculture industry.

Health & Welfare
Microsatellites: Applications in aquaculture
Microsatellites are useful for molecular tagging of individuals and molecular dissection of complex traits such as growth, temperature tolerance and disease resistance.

Health & Welfare
Genome editing: Potential to improve aquaculture breeding, production, Part 1
Due to high fecundity and external fertilization, most aquaculture species are amenable to genetic improvement technologies, including genome editing.


