Detection of WSSV critical for shrimp-farming facilities
Causing high mortality in penaeid shrimp, white spot syndrome (WSS) has devastated the shrimp aquaculture industry worldwide. WSS is characterized by white spots of 0.5 to 2.0 mm in diameter on the shrimp cuticle, which may also become loose and reddish. Affected animals exhibit lethargy accompanied by a sudden reduction in food consumption.
The disease is caused by white spot syndrome virus (WSSV), a large double-stranded DNA virus. WSSV was first reported in Taiwan in 1992 and in East and South Asia in following years. It appeared in North, Central and South America by 2000. More than two decades after the first report, Australia is the only continent to have avoided WSSV epidemics.
WSSV has a broad host range within aquatic crustaceans. More than 93 host species have been reported, including penaeid shrimp, freshwater shrimp, crayfish, crabs, lobsters and oysters.
In most cultured penaeid shrimp, transmission of the virus can occur vertically from infected parents or horizontally by direct contact with infected individuals or virus particles in the water, or by ingestion of infected tissues. In addition, planktonic organisms, rotifers and polychaetes are potential vectors of WSSV.
Health management
Without vaccines, biosecurity plays a very important role in the health management of cultured shrimp. Biosecurity aims to prevent pathogens from entering the culture environment and hence reduce the economic impacts of diseases.
Biosecurity measures include screening for seed of good quality, establishment and maintenance of a WSSV-free facility and routine surveillance of the shrimp population for early detection and quarantine. Sensitive and specific diagnostic tools capable of detecting low levels of WSSV are essential for these measures.
The advances of biotechnology in recent decades have created molecular detection methods that differ in sensitivity, specificity, sample throughput, costs of devices and reagents, risk of cross contamination and ease of data interpretation. Various commercial WSSV detection kits are available to meet the needs of different shrimp-farming facilities.
Polymerase chain reaction
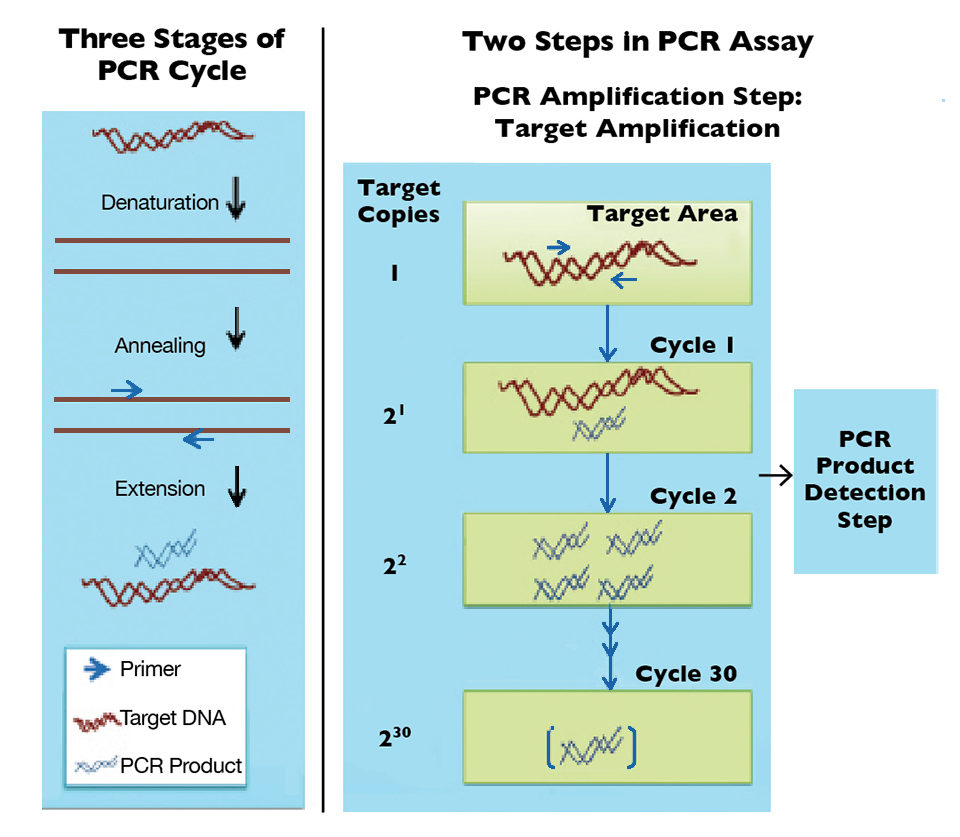
Offering high sensitivity and specificity, polymerase chain reaction (PCR) assays are currently the most accepted molecular method for WSSV detection. PCR assays generally involve two steps: target amplification by PCR and product detection. PCR relies on a heat-resistant DNA polymerase enzyme to copy double-stranded DNA templates from short target-specific primers.
Basically, PCR cycles from strand separation or denaturation of target DNA to annealing of primers for target DNA and extension from primers to make copies of the target DNA, resulting in a twofold increase after each cycle (Figure 1). PCR primers identify the target specifically by recognizing novel signatures found within the genetic materials of target organism. Ideally, only the desired product is produced in large amounts from the sample at the end of all cycles.
After target amplification, the reactions need to be processed manually for signal detection in some assays. In addition, these assays normally require hands-on time to assemble the reactions before the target amplification reaction. Because these methods involve high sensitivity, proper laboratory operation protocols need to be implemented to minimize risks of cross contamination, which can lead to false positive or negative results.
Commercial assays
Commercial WSSV assays are based on different PCR methods with minor variations in the reaction device and mechanisms of signal generation and detection. These methods include “one-step” PCR, nested PCR, real-time quantitative PCR (qPCR) and insulated isothermal PCR (iiPCR). The properties of these methods are summarized in Table 1.
Lee, Summary of PCR methods, Table 1
| PCR | Nested PCR | Real-Time qPCR | iiPCR | |
|---|---|---|---|---|
| Specificity | +++ | +++ | +++ | +++ |
| Sensitivity | + | +++ | +++ | +++ |
| Quantitation | Semi-quantitation possible | Semi-quantitation possible | Yes | No |
| Product detection | Visual | Visual | Automatic reading | Automatic reading |
| Hands-on time | ++ | +++ | + | + |
| Throughput | ++ | ++ | +++ | + |
| Post-amplification processing | Yes | Yes | No | No |
| Cross contamination | +++ | +++ | + | + |
| Reaction time | ++ | +++ | + | + |
| Cost of reagents | + | + | +++ | ++ |
| Cost of device | + | + | +++ | + |
| Device portable | No | No | No | Yes |
One-step PCR assays
Conventional one-step PCR assays include only basic amplification of target DNA, leading to sensitivity lower than that of the other PCR formats. Product detection relies mainly on gel analysis, which involves the separation of products through a gel of certain pore sizes in an electronic field (gel electrophoresis) and staining and visual detection of product bands on the gel with the help of a fluorescent dye.
Target amplification and product detection usually take two to three hours to complete. Besides being labor-intensive and time-consuming, these procedures can bring risks of cross contaminating amplicons into future assays, leading to false positive results. However, assays based on this format are generally cheaper than the other PCR assays.
Nested PCR assays
To improve the sensitivity and specificity of one-step PCR, products of the first assay can be amplified further in a nested, second round of PCR. This type of assay requires more hands-on time than the other methods because it includes two assay assembly steps and usually requires gel analysis of reaction products.
Target amplification and product detection (gel analysis) usually take three to four hours. However, the possibility of semi-quantitative analysis of WSSV provides important information about viral loads in shrimp samples to follow the progression of the disease. This is currently the method most recognized and accepted by the shrimp culture industry because of its satisfactory sensitivity, acceptable costs and reasonable sample throughput. Among the commercial WSSV kits, the IQ2000 WSSV Detection and Prevention System has been certified by the World Organisation for Animal Health (OIE) as fit for the diagnosis of white spot disease in crustaceans.
Real-time quantitative PCR assays
In real-time qPCR, integration of a fluorescent dye/probe in the PCR reaction and optical detection module in the PCR apparatus enable easy qualitative and/or quantitative detection of amplicon production. However, real-time qPCR requires a sophisticated and expensive device for reaction and signal detection.
Inclusion of fluorescent dye-linked probes makes these assays more sensitive, yet more costly than single-step PCR. The advantages of real-time qPCR include the fact that results can be obtained without opening the reaction tubes, greatly reducing cross-contamination risks that come with gel analysis of PCR products. Furthermore, real-time qPCR usually takes one to two hours to complete and allows quantitative detection of WSSV to assist assessment of the severity of disease outbreaks. Capable of high-throughput analysis, real-time qPCR can fulfill the need of large-scale farms to screen large numbers of samples in a PCR lab.
Insulated isothermal PCR assays
The iiPCR assay is based on a new PCR format that requires only a simple heat source to drive the PCR cycles. It works in a portable nucleic acid analyzer, designed specifically for iiPCR with capability to detect products automatically and make pond-side detection of WSSV possible. It is similar to real-time qPCR in providing automatically processed results of high sensitivity without any post-reaction steps.
Automatic reading of product signals makes the assay reliable and user-friendly, and reduces human errors and discrepancies in data interpretation. iiPCR assays can finish in approximately 1.5 hours. The costs of reagents and devices for iiPCR assays are lower than those for real-time qPCR assays. However, iiPCR assays do not provide quantitative scores, and the devices can process 8 reactions/run. The only iiPCR-based assay for WSSV, I.Q. Plus WSSV Kit with POCKIT System, was recently certified by the OIE as fit for the diagnosis of white spot syndrome.
Perspectives
Sensitive and specific detection of WSSV at appropriate check points is critical for all shrimp-farming facilities. Depending on the budget, location and resources of the facility, different PCR assays can be selected to fulfill the needs for WSSV detection.
For large-scale farms, which could hire a specialist and afford real-time PCR reagents and machines, real-time qPCR assays are recommended because of their high-throughput capacity and short hands-on time. Nested PCR assays provide satisfying results for facilities with limited budgets. On the other hand, the portable and user-friendly iiPCR assays are suitable for small-scale farms, where only a small number of samples need to be tested at each check point.
(Editor’s Note: This article was originally published in the September/October 2013 print edition of the Global Aquaculture Advocate.)
Authors
-
Dr. Pei-Yu Alison Lee
Vice President, Research and Development
GeneReach Biotechnology Corp.
No.19, Keyuan 2nd Road
Central Taiwan Science Park
Taichung, 407 Taiwan[109,111,99,46,104,99,101,116,111,105,98,104,99,97,101,114,101,110,101,103,64,57,50,51,117,121,105,101,112]
-
Dr. Ping-Hua Teng
Vice President, Technical Service
GeneReach Biotechnology Corp.
No.19, Keyuan 2nd Road
Central Taiwan Science Park
Taichung, 407 Taiwan -
Simon Chung
Vice President, Sales
GeneReach Biotechnology Corp.
No.19, Keyuan 2nd Road
Central Taiwan Science Park
Taichung, 407 Taiwan
Tagged With
Related Posts

Health & Welfare
A comprehensive look at the Proficiency Test for farmed shrimp
The University of Arizona Aquaculture Pathology Laboratory has carried out the Proficiency Test (PT) since 2005, with 300-plus diagnostic laboratories participating while improving their capabilities in the diagnosis of several shrimp pathogens.

Health & Welfare
CENIACUA develops WSSV-resistant shrimp in Colombia
To combat white spot syndrome virus (WSSV) in white shrimp, Corporación Centro de Investigación de la Acuacultura de Colombia (CENIACUA) initiated a selective-breeding program to develop resistance in shrimp.
Health & Welfare
Double-stranded RNA against WSSV genes provides antiviral protection in shrimp
Silencing genes in white spot syndrome virus (WSSV) with critical roles in replication could provide a strong antiviral effect and thus reduce shrimp mortality. The authors therefore established a study to evaluate the antiviral efficacy of double-stranded (ds)RNA against non-structural WSSV genes.

Health & Welfare
Mineral extract reduces EMS, WSSV impacts in Pacific white shrimp
The authors recently performed a study to evaluate immunity to white spot syndrome virus and virulent Vibrio parahaemolyticus in shrimp treated with a new mineral extract additive in pelletized feeds.


