Scombrotoxin poisoning accounts for large share of U.S. seafood-related illnesses

Histamine and other biogenic amines are present in various amounts in many foods. Fresh fish at harvest are virtually free of histamine, but post-harvest conditions that allow the growth of spoilage bacteria can result in histamine formation.
Histamine fish poisoning
Histamine (or scombrotoxin) fish poisoning occurs worldwide through ingestion of fish containing hazardous levels of histamine and other biogenic amines such as putrescine and cadaverine. In the United States, this poisoning accounted for 7.5 percent of all foodborne outbreaks and 38 percent of all seafood-related illnesses reported from 1990 to 2003. From 1998 to 2007, 321 outbreaks involving 1,328 cases with 58 hospitalizations and no deaths were reported to the Centers for Disease Control and Prevention.
Human illness occurs rapidly after ingestion of fish with elevated histamine levels and lasts from several minutes to a few hours. Symptoms include allergic-like responses such as headache, dizziness, swelling of the tongue, nausea, vomiting, diarrhea and stomach pain. Histamine fish poisoning is usually self-limiting, and recovery is complete.
Histamine is produced by certain spoilage organisms through action of the enzyme histidine decarboxylase (HDC), which converts the amino acid histidine to histamine. There are two classes of HDC enzymes. One class is found in eukaryotic cells and Gram-negative bacteria. The second group is found in Gram-positive bacteria.
Both groups have different coenzymes associated with them. Gram-positive histamine-producing bacteria are more commonly associated with fermented products like salami, cheese, sauerkraut and wine, while Gram-negative histamine-producing bacteria are more common in fish. A wide range of Gram-negative bacteria can produce histamine in fish, but the major types are mesophilic enteric and marine bacteria.
The U.S. Food and Drug Administration considers fish with over 50 ppm histamine decomposed, and fish containing greater than 500 ppm histamine are a human health hazard. Therefore, high-histamine-producing bacteria pose a greater risk for human health than low-histamine producers.
Analysis
Methods for the analysis of histamine in fish include qualitative and quantitative immuno-based test kits, as well as high-performance liquid chromatographic methods. A better understanding of the relationships between the presence of histamine-producing bacteria and formation of histamine in fish is needed in order to develop effective mitigation strategies.
Prompt analysis for histamine-producing bacteria can help identify conditions that lead to histamine production and histamine fish poisoning. Therefore, reliable, rapid and accurate methods for the detection of histamine-producing bacteria in fish are needed.
Traditional detection methods
Histamine-producing bacteria are traditionally detected by differential culture-based methods using agar or broth (Table 1). Although these methods are easy to use and inexpensive, they have several disadvantages. For example, the low (5.3) pH of the medium can inhibit the growth of some histamine-producing bacteria. And some bacterial strains, such as Photobacterium damselae and Hafnia alvei, do not grow well at low pH. However, increasing pH can increase the risk of false positive reactions.
Butler, Comparison of detection methods, Table1
| Method | False Positive | Direction | Results | Time |
|---|---|---|---|---|
| Differential agar | 15-75% | High- and low-HPB | Qualitative | 2 days |
| Differential broth | – | High- and low- HPB | Qualitative/quantitative | 2 days |
| Colony lift hybridization | 0 | High-HPB | Quantitative | 2 days |
| Traditional PCR | 0 | High-HPB | Qualitative | 1-2 days |
| Real-time PCR | 0 | High-HPB | Quantitiative | 1 hour |
Various authors have reported 15 to 72 percent false positive results due to other bacterial metabolic processes that increase the pH of the medium and lead to a color change similar to that resulting from histamine production. For these reasons, differential culture-based methods are considered best suited in screening fish for the presence of histamine-producing bacteria with subsequent confirmation of the production of histamine by chemical methods.
Another detection technique employs physicochemical analysis. One potentiometric-based method, which detects histamine-producing bacteria in fish by measuring increased conductance in culture media, can be used to screen bacterial isolates. It is highly effective for the detection of high-histamine-producing bacteria but is not suitable for detection of low histamine producers.
Molecular techniques
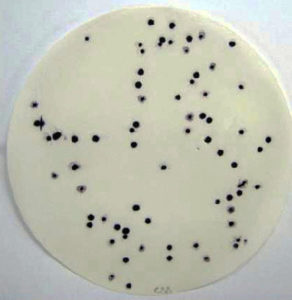
producing bacteria in fish.
Recent research has focused on the development of molecular-based techniques targeting the HDC gene from Gram-negative histamine-producing bacteria. Several investigators have used nucleic acid amplification methods targeting various fragments of the HDC gene. A polymerase chain reaction (PCR) method that amplifies the variable region of the 16S rDNA gene from the high-histamine producer (Morganella morganii) was developed for its detection. Methods such as these can be used to confirm bacterial cultures that screen positive on Niven’s medium, but they do not eliminate the need to verify histamine production by chemical analysis.
Although molecular-based methods result in fewer false positive results than the culture-based methods, they appear to detect only the high-histamine-producing bacteria. The authors compared culture-based, potentiometric and traditional PCR methods for the detection of histamine-producing bacteria and found that while the Niven’s method detected both high- and low-histamine-producing strains, it gave a high number (38 percent) of false positive results. The potentiometric and PCR-based methods produced no false positives for high-histamine-producing strains, but failed to detect low histamine producers.
Failure to detect low-histamine-producing bacteria by the molecular-based methods may have several explanations. Firstly, the specific gene in the low histamine producers is not yet known. It is possible that these bacteria do not carry the HDC gene and produce histamine by some other pathway. In fact, other amino acid decarboxylases are known to decarboxylate histidine in addition to their natural substrates.
Secondly, the HDC gene in these bacteria may be plasmid associated and not detected by the molecular method. It could also be lost during subculturing of the strains.
Combined culture, molecular techniques
To overcome some limitations inherent to molecular techniques, the authors combined culture-based methods with nucleic acid hybridization.
Colony lift hybridization is one technique that is uniquely suited to situations where the performance of selective or differential media is less than perfect, as it provides more accurate quantitative results because the target organism can be confirmed without the need for subculturing. The authors therefore developed digoxigenin-labeled probes for the detection and quantification of histamine-producing bacteria using colony hybridization. This new method targets high-histamine-producing strains of bacteria based on the specificity of probes targeting the HDC gene of these organisms.
A probe mix created by PCR amplification and labeling of the HDC gene of four high-histamine-producing strains (M. morgannii, R. planticola, E. aerogenes and P. damselae) performed extremely well when applied to strains producing over 1,000 ppm histamine. The study showed 100 percent specificity for high-histamine-producing strains using the probe mix against a culture library of 152 Gram-negative histamine- and non-histamine-producing bacteria.
The method did not detect or discriminate between low- and non-histamine-producing bacteria. However, the study provided evidence that colony lift hybridization can be used to detect and quantify high-histamine-producing bacteria.
More recently, a real-time PCR method for the detection of histamine-producing bacteria was developed by researchers at the U.S. Food and Drug Administration. Real-time PCR is considerably less time-consuming than traditional PCR, with results normally obtained within an hour. In addition to template amplification, an internal control nucleic acid molecule can be incorporated into the assay to prevent false negative results.
Perspectives
The development of reliable, rapid and accurate methods for detection of histamine-producing bacteria in fish remains a goal of research. Important steps have been taken to develop faster and more reliable molecular-based methods, but additional research is needed for the detection of low-histamine-producing bacteria.
Prompt analysis for histamine-producing bacteria can be a useful tool to identify conditions that lead to histamine production and histamine fish poisoning. As more-comprehensive detection methods are developed, they can be validated against known histamine-producing organisms to determine the risk for histamine fish poisoning.
(Editor’s Note: This article was originally published in the September/October 2010 print edition of the Global Aquaculture Advocate.)
Now that you've reached the end of the article ...
… please consider supporting GSA’s mission to advance responsible seafood practices through education, advocacy and third-party assurances. The Advocate aims to document the evolution of responsible seafood practices and share the expansive knowledge of our vast network of contributors.
By becoming a Global Seafood Alliance member, you’re ensuring that all of the pre-competitive work we do through member benefits, resources and events can continue. Individual membership costs just $50 a year.
Not a GSA member? Join us.
Authors
-
Kristin Björnsdóttir-Butler, Ph.D.
Post-Doctoral Fellow
USFDA Gulf Coast Seafood Laboratory
1 Iberville Drive
Dauphin Island, Alabama 36528-01581 USA[118,111,103,46,115,104,104,46,97,100,102,64,114,101,108,116,117,98,46,110,105,116,115,105,114,107]
-
David P. Green, Ph.D.
Professor
Department of Food, Bioprocessing and Nutrition Sciences
North Carolina State University
Center for Marine Sciences and Technology
Morehead City, North Carolina, USA
Related Posts

Health & Welfare
AHPN inferences based on behavior of vibrio bacteria
Vibrio parahaemolyticus, a strain of which is the cause of acute hepatopancreatic necrosis (AHPN), has both virulent and benign strains. This strain colonizes the stomachs of shrimp by the formation of a biofilm, which protects it from antibiotics and other potential treatments.

Health & Welfare
A holistic management approach to EMS
Early Mortality Syndrome has devastated farmed shrimp in Asia and Latin America. With better understanding of the pathogen and the development and improvement of novel strategies, shrimp farmers are now able to better manage the disease.

Health & Welfare
A comprehensive look at the Proficiency Test for farmed shrimp
The University of Arizona Aquaculture Pathology Laboratory has carried out the Proficiency Test (PT) since 2005, with 300-plus diagnostic laboratories participating while improving their capabilities in the diagnosis of several shrimp pathogens.

Intelligence
Post-harvest quality of freshwater prawns, part 2
Freshwater shrimp can contain pathogenic bacteria that cause illness unless care is exercised by producers, retailers and consumers. Many of the human pathogens can survive frozen storage, but are killed or inactivated by thermal processes.


