Study results useful as initial screening method

The protozoan Paramoeba sp. (synonymous to Neoparamoeba sp.) is ubiquitous and normally free-living in marine waters, and it is the most pathogenic amoeboid protozoan parasite in cultured fish. The species P. perurans colonizes gills and results in amoebic gill disease (AGD) in farm-raised salmonids and most notably affects the Tasmanian Atlantic salmon industry, with typical losses of 10 to 20 percent of production. So far, AGD has been recorded in 15 finfish species of 11 different genera, and in fish, P. perurans infection leads to proliferative cell reactions in the gills, including epithelial hyperplasia, hypertrophy, edema and inter-lamellar vesicle formation, which grossly appear as white mucoid spots and plaques on the gill surface.
Gross pathological assessment of the gill has been widely used for AGD diagnosis, but a subsequent histological examination is recommended to improve diagnostic accuracy. Additionally, molecular diagnostic methods based on conventional PCR assay targeting the small subunit rRNA (SSU rRNA) sequence are available for the diagnosis of AGD.
Various species of Paramoeba have also been reported in some crustacean and echinoderm species, such as lobsters, crabs and sea urchins. In these hosts, severe infections were associated with the death of the host, resulting in significant economic losses. The infections were also reported in bivalve mollusks, such as mussels, but these were considered more likely to be potential environmental reservoirs for this protozoan.
This article – adapted and summarized from the original publication – is the first report of this amoeboid protozoan parasite in Pacific white shrimp (Litopenaeus vannamei). This work was supported by the National Research Foundation of Korea (NRF) grant funded by the Korea government (MSIT) (grant number NRF-2018R1C1B5086350).
Study setup
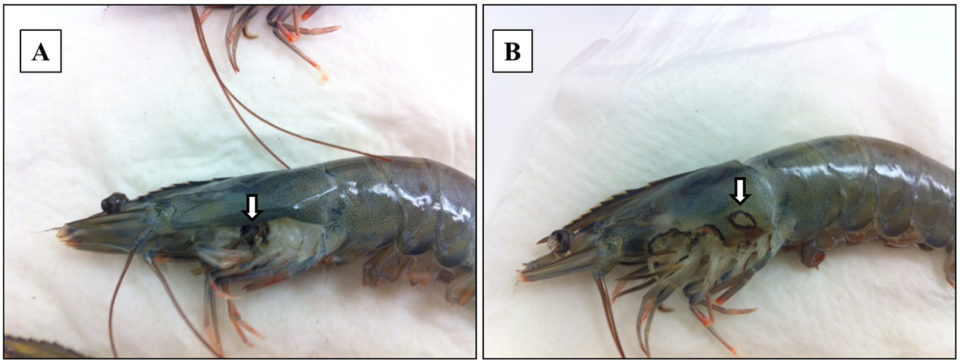
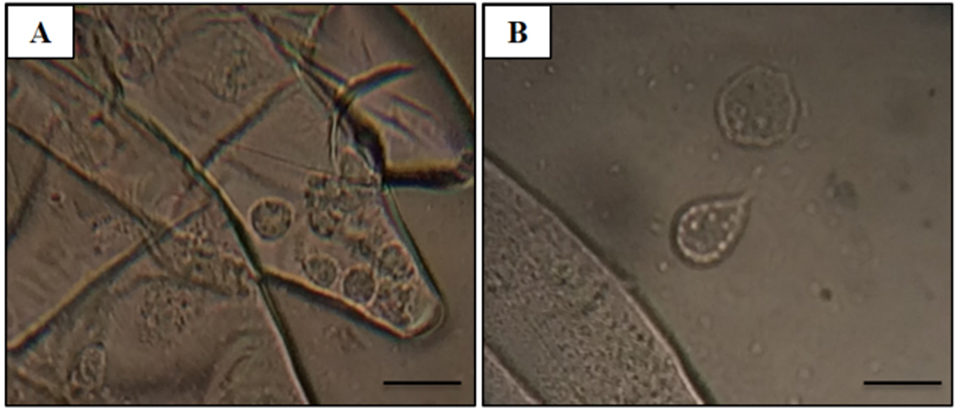
Adult Pacific shrimp (avg. weight 30 grams) from an anonymous shrimp hatchery in North America showed reduced appetite, lethargic, and respiratory distress. Cumulative mortality was 62.75 percent at 120 days after stocking, and diseased shrimp had black gills, missing lamellae and eroded carapace (see picture above). From these data, we suspected a fungal infection caused by Fusariumspecies, which has been previously reported as the causative agent of black gill disease in crustaceans, like prawns, shrimp and lobsters. However, amoebic protozoan parasites were observed by direct observation of a wet mount (Fig. 1). Moribund shrimp samples were collected at irregular intervals, fixed in Davidson’s AFA fixative or 95 percent ethanol, and further examined by histopathology, PCR and ISH.
For additional, detailed information on shrimp sampling, histopathology examination and sequence analysis, PCR assay and in situ hybridization (ISH), please consult the original publication.
Results and discussion: Histopathology examination (H&E)
The most significant finding in the microscopic examination of the studied samples was the presence of an amoebic parasite. Infestation by this parasite was found mainly in the gills, with severity grade G4 (Figs. 2A–C), and extensive hyperplasia, bridging of lamellae, and forming of inter-lamellar spaces were observed. According to the farmer, lethargy, anoxia, and, eventually, death has been seen in infected shrimp, and these might be associated with the damage to the gills (respiratory organs). Fig. 2A shows the amoeba nucleus with an amphiphilic core surrounded by an irregular basophilic ring and the parasome with eosinophilic cytoplasm and vacuoles, indicating that the amoeba is closely related to the members of the order Dactylopodida, and maybe the genus Paramoeba, according to Sühnel et al. (2014).
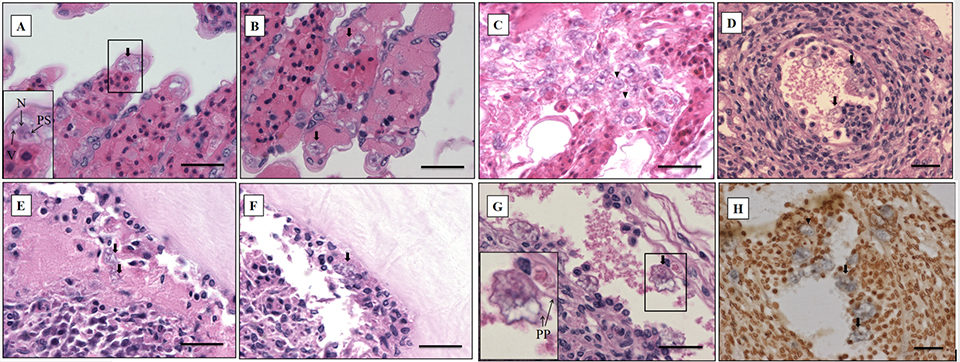
In addition, flattened and irregular-shaped trophozoite forms with pseudopodia radiation (approximately 25 to 30 μm in diameter) were predominantly observed. The presence of digitate pseudopodia, lack of cysts (less than 10 μm in diameter), and marine habitat also identify the amoeba as the genus Paramoeba, according to Kent et al. (1988). These amoebic parasites were also observed in other organs, including the antennal gland, lymphoid organ, cuticular epithelium, and the subcutis of appendages and connective tissue surrounding the ventral nerve cord of the tested samples, with severity grade G1 to G4 (Figs. 2D–G).
PCR assay and sequence analysis
Strong amplicons (an amplicon is a piece of DNA or RNA that is the source and/or product of amplification or replication events) were detected in the tested shrimp (five representative samples), by the PCR assay targeting the SSU rRNA sequence (Fig. 3). Results showed that the nucleotide sequence was 100 percent identical to the sequences of Paramoeba sp. (Neoparamoeba sp.) from several marine crustacean and echinoderm species. And there was no cross-reaction with genomic DNA from L. vannamei shrimp and other shrimp (P. monodon, P. indicus, L. stylirostris, and M. rosenbergii), polychaetes, squid and Artemia spp., or to other shrimp parasites by the specificity test.
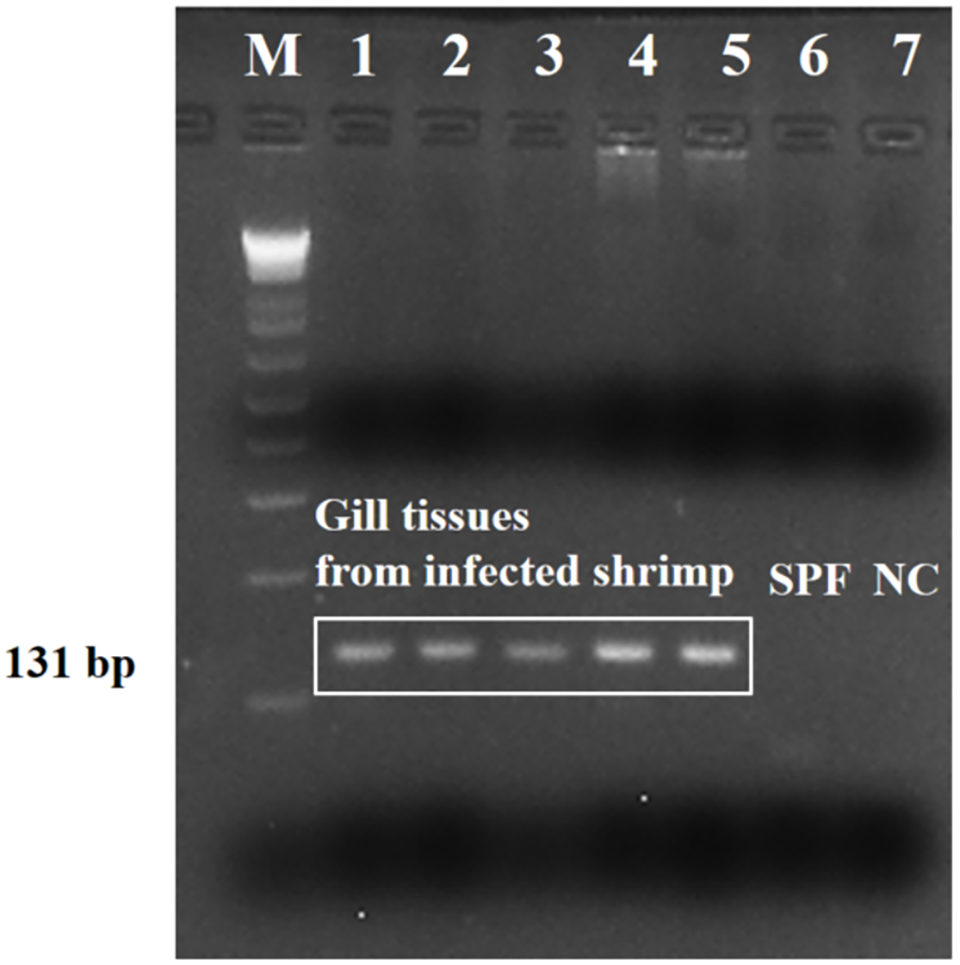
We also conducted a phylogenetic study to investigate the relationship between identified Paramoeba-like sp. from shrimp and other amoeba species. In the resulting phylogenetic tree, the SSU rRNA sequence from Paramoeba-like sp. was grouped together with sequences from the order Dactylopodida, especially the family Paramoebidae.
In situ hybridization
The ISH method can also be used to determine the etiological agents of protozoan infection. For ISH, the Paramoeba-like gene probe was generated from the SSU rRNA sequence of the amoeba infecting shrimp, and this hybridized with the amoebic cells in the representative sample (Fig. 2H), corresponding to the histopathology results. The probe appears to be highly specific, and no reaction was seen in any of the tissues prepared from SPF shrimp (data not shown).
So far, the amoeboid protozoan parasite Paramoeba sp. has been reported in several marine crustaceans, including American lobster and crab, but not in cultured shrimp. This study is the first report of infection by the amoebic protozoan parasite Paramoeba-like sp., in the gill of L. vannamei cultured in an anonymous shrimp hatchery in North America. It is most likely that the amoeba infection was due to stress factors, such as increased water temperature and/or high salinities, combined with high densities of animals in ponds, which provide an advantage to this protozoan naturally present in the marine environment. According to farmers, the salinity had dropped from 34 to 10 ppt due mechanical equipment issues, so it is likely that the resulting stress had triggered the parasite infestation in shrimp.
Perspectives
Shrimp infected with the amoeba showed reduced appetite, lethargy, respiratory distress, eroded carapaces and blackened gills. Under the microscope, histopathological characteristics included the typical morphology of amoebic protozoa displaying granular endoplasm with vacuoles, nucleus, parasome and ectoplasm with pseudopodia. These features suggest that the parasite is related to the order Dactylopodida, probably a Paramoeba species and this was confirmed by the SSU rRNA sequence analysis, PCR assay and in situ hybridization (ISH).
In shrimp, the amoeba infection has resulted in significant mortality and associated economic losses. Therefore, the ISH and PCR diagnostic assays developed in this study can provide informative data for shrimp farmers and can be used as the initial screening methods for amoeboid protozoan in shrimp.
This study provides informative data to shrimp producers and helps farmers monitor for amoeba infections in shrimp farms. In the examined shrimp, the parasites have histological characteristics of Paramoeba sp., but no PCR bands were detected for the primers generated from P. perurans, P. pemaquidensis, or P. branchiphila spp. Therefore, we assume that it could be a new Paramoeba species infecting shrimp. Additional work is needed to diagnose the species and develop the species-specific diagnostic methods for amoeba infecting farmed shrimp.
References available from first author or the original publication.
Now that you've reached the end of the article ...
… please consider supporting GSA’s mission to advance responsible seafood practices through education, advocacy and third-party assurances. The Advocate aims to document the evolution of responsible seafood practices and share the expansive knowledge of our vast network of contributors.
By becoming a Global Seafood Alliance member, you’re ensuring that all of the pre-competitive work we do through member benefits, resources and events can continue. Individual membership costs just $50 a year.
Not a GSA member? Join us.
Authors
-
Jee Eun Han, DVM, Ph.D.
Laboratory of Aquatic Biomedicine
College of Veterinary Medicine
Kyungpook National University
Daegu 41566, Korea[109,111,99,46,108,105,97,109,103,64,51,50,50,49,101,106,110,97,104]
-
Ji Hyung Kim, Ph.D.
Senior researcher
Infectious Disease Research Center
Korea Research Institute of Bioscience & Biotechnology (KRIBB)
125 Gwahak-ro, Yuseong-gu, Daejeon 34141, Republic of Korea -
Kyeong Yeon Kim
Laboratory of Aquatic Biomedicine
College of Veterinary Medicine
Kyungpook National University
Daegu 41566, Korea -
Young Seo Lee
Laboratory of Aquatic Biomedicine
College of Veterinary Medicine
Kyungpook National University
Daegu 41566, Korea
Tagged With
Related Posts

Health & Welfare
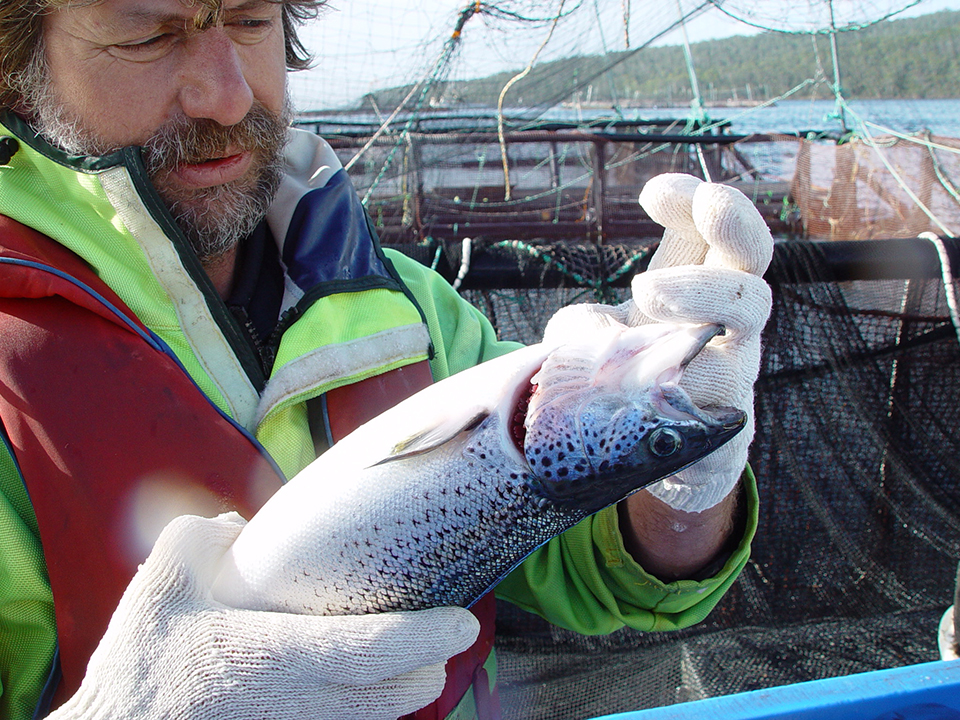
Developing a breeding program for Tasmanian salmon
A research partnership focused on Atlantic salmon selective breeding aims to improve growth and resistance to amoebic gill disease in Tasmanian salmon.

Innovation & Investment
Aquaculture America 2017: Communication key to the future
This year’s Aquaculture America in San Antonio, Texas, provided significant learning and networking opportunities. It successfully brought together 14 U.S. aquaculture organizations and more than 1,600 participants from Europe, Asia, Africa and Australia.

Health & Welfare
Biofloc technology: Possible prevention for shrimp diseases
Facing emerging viral problems and rising energy costs, the use of biofloc technology in biosecure systems offers an answer for sustainable shrimp aquaculture. The main attributes of biofloc systems in reducing disease risk include the fact that low water exchange improves pathogen exclusion.

Health & Welfare
EHP a risk factor for other shrimp diseases
Laboratory challenges and a case-control study were used to determine the effects of EHP infection on two Vibrio diseases: acute hepatopancreatic necrosis disease (AHPND) and septic hepatopancreatic necrosis (SHPN).



