Bacteri, yeasts isolated from native microbiota of wild shrimp
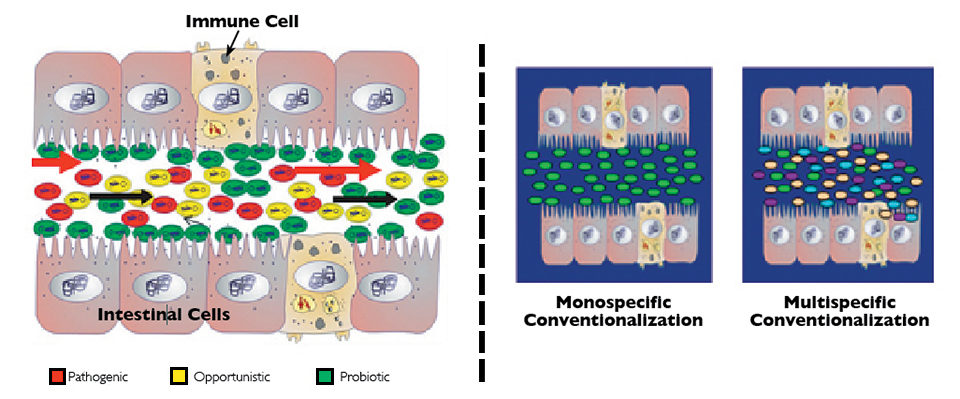
The successful culture of shrimp at early stages may essentially depend on digestive tract colonization by beneficial microorganisms and exclusion of pathogenic ones. Native microbiota have been shown essential for correct development and function of the digestive tract and its associated immune system.
However, aquaculture production practices and systems do not always permit the process of digestive tract colonization with native microbiota. Water, eggs and nauplii are disinfected, but later contaminated with opportunistic bacteria present in the culture systems (microalgae, nauplii, zooplankton, feed) and not related to the native microbiota in cultivated animals.
With the support of FINCYT, domestication of native microbiota in white leg shrimp (Litopenaeus vannamei) has been investigated by the authors’ teams at Marinazul in Peru and Concepto Azul in Ecuador. Microorganisms isolated from different parts of the shrimp digestive tract have been cultivated, molecularly characterized and evaluated according to a process known as “conventionalization.” Conventionalized larvae – animals with global or partial native microbiota – showed better growth and/or survival in comparison to the normal hatchery-produced larvae.
Study setup
Bacteria and yeasts were isolated from the native microbiota of wild shrimp at larval, juvenile and adult life stages. Characterizations of the isolated bacteria and yeasts were based on metabolic assays of amylase and protease, antagonism tests with pathogenic bacteria and molecular criteria related to amplification and sequencing of 16S rDNA, rpoB and lactonase genes.
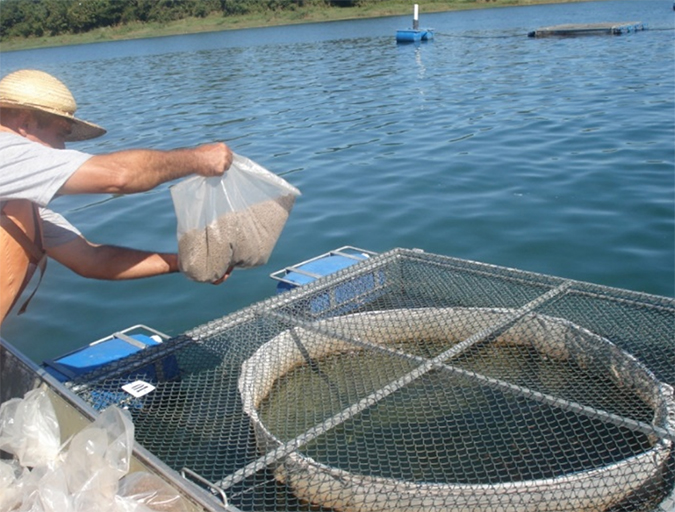
Mono- and multispecific conventionalization processes were performed by placing the shrimp in the respective suspensions of microorganisms. As a preliminary step, the authors established protocols for egg and larva disinfection before inoculation of microbial strains. Just after spawning, shrimp eggs were disinfected with formalin. A second disinfection process was applied to stage IV shrimp nauplii just before stocking them in sterilized seawater.
The selected microorganisms were administered directly into the culture water before and after the mouth-opening period of the shrimp. One inoculation was given every other day until the PL12 stage. Effective digestive tract colonization could be qualitatively and quantitatively monitored using laser-scanning confocal microscopy and real-time polymerase chain reaction testing.
At the end of the experimental period, growth and survival rates were estimated. In some cases, experimental infections with pathogenic bacteria were performed to estimate the protection efficiency of native microorganisms in conventionalized animals.
Results
Three specific collections of native microbiota microorganisms were established and molecularly characterized, corresponding to yeasts (Saccharomyces, Moeszyomyces, Candida), Gram-negative and Gram-positive bacteria belonging to the genera Vibrio, Aeromonas, Acinetobacter, Oxalobacter, Photobacterium, Rhodobacter, Psychrobacter, Psychromonas, Pseudomonas, Shewanella, Proteus, Enterococcus, Micrococcus, Microbacterium, Bacillus and Lysinibacillus, Exiguobacterium and a collection of unknown bacterial strains.
Along the hepatopancreas, midgut and hindgut of the shrimp digestive tract, Gram-positive proteobacteria were predominant, with a majority of Vibrio species followed by Photobacterium, Psychobacter and Shewanella. Rhodobacter was only present in adult and juveniles. Gram-positive bacteria were principally represented by Bacillus species, followed by Micrococcus, Exiguobacterium and Enterococcus in adult and juveniles.
Low bacteria diversity was observed in shrimp larvae, where Vibrio, Psychobacter and Bacillus species were mostly present. Among Vibrios, the species alginolyticus was dominant (Table 1).
Emmerik, Cultivable bacterial distribution, Table 1
| Bacteria Species | Adult Shrimp | Juvenile Shrimp | Postlarvae |
|---|---|---|---|
| Vibrio | 62.35% | 32.0% | 42.0% |
| Photobacterium | 17.65% | 6.0% | – |
| Shewanella | 4.71% | – | – |
| Bacillus | 3.53% | 21.0% | 16.0% |
| Aeromonas | 3.53% | 8.0% | 5.0% |
| Rhodobacter | 2.35% | 2.0% | – |
| Psychrobacter | 2.35% | 6.0% | 37.0% |
| Micrococcus | 1.18% | – | – |
| Leptolyngbya | 1.18% | 2.0% | – |
| Exiguobacterium | 1.18% | 13.0% | – |
| Proteus | – | 3.0% | – |
| Psychromonas | – | 3.0% | – |
| Microbacterium | – | 2.0% | – |
| Enterococcus | – | 2.0% | – |
The selection of bacteria with metabolic characters potentially interesting for probiotic effects (protease, amylase and lactonase) was conceptualized to constitute several mixes for conventionalization.
Quorum quenching
Vibrio bacteria possess an elaborate communication system known as “quorum sensing,” which relies on extracellular production of lactone. High concentrations of lactone serve to induce such gene functions as bioluminescence, biofilm formation and toxin production. Nevertheless, some vibrio-antagonistic bacteria developed a system able to inhibit quorum sensing known as “quorum quenching” through degradation of lactone by the enzyme lactonase.
Most lactonase-producing bacteria are Bacillus or Schewanella. Identification of such bacteria was performed using polymerase chain reaction testing in order to detect the presence of the lactonase gene sequence in the bacterial genome.
The efficiency of several strains considered potential probiotics able to inhibit three pathogenic vibrios was experimentally tested. Administration of a lactonase-producing bacteria mix to shrimp postlarvae experimentally infected by pathogenic vibrio was efficient and showed up to 10 times higher survival at three days post-infection than in infected vibrio without administration of probiotics.
Conventionalization effects
The use of strains able to produce extracellular amylase and protease could be very important to improve digestion and assimilation of balanced diets. Recent studies have shown that the shrimp domestication process resulted in a loss in capacity to process carbohydrates and favored selection of shrimp with greater capacity to digest proteins as an energy source.
Wild shrimp are characterized by their primary capacity to digest glucides, a “cheaper” energy source than protein. As a consequence, the selection and application of probiotics represent necessary tools to compensate for this drawback of domestication in feeds formulated with lower protein content without negative impacts on growth and survival rates. Moreover, simplification of ingested feeds by bacterial enzymatic activity should allow fast, optimized nutrient assimilation by shrimp digestive systems.
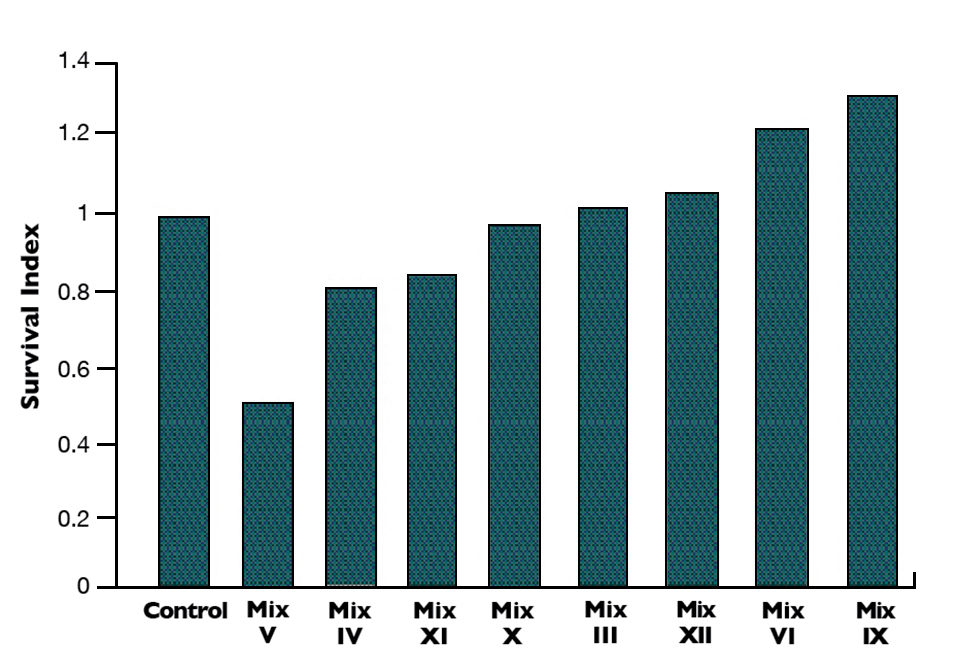
With the first eight bacterial mixes tested in triplicate, four mixes gave better survival in Litopenaeus vannamei than the control provided (Fig. 1). Results showed a 10 to 30 percent increase in survival with three combinations of Bacillus and Vibrio alginolyticus native strains. Microbial mixes IV and V, made mainly with Gram-negative pathogenic/opportunistic bacteria, showed strong negative impacts on survival.
Perspectives
It is important to consider that every animal and plant species has performed a coevolution process with its native microbiota. Aquaculture will have to consider such processes for present and future culture species. Because of the recent progress in molecular microbiology and confocal microscopy, applied research in aquaculture gnotobiology has been made possible, as showed by the present work with some model invertebrate and vertebrate species.
(Editor’s Note: This article was originally published in the November/December 2010 print edition of the Global Aquaculture Advocate.)
Now that you've reached the end of the article ...
… please consider supporting GSA’s mission to advance responsible seafood practices through education, advocacy and third-party assurances. The Advocate aims to document the evolution of responsible seafood practices and share the expansive knowledge of our vast network of contributors.
By becoming a Global Seafood Alliance member, you’re ensuring that all of the pre-competitive work we do through member benefits, resources and events can continue. Individual membership costs just $50 a year.
Not a GSA member? Join us.
Authors
-
Motte Emmerik
Concepto Azul S.A.
P.O. Box 0902-142A
Guayaquil, Ecuador[109,111,99,46,108,105,97,109,103,64,107,105,114,101,109,109,101,46,101,116,116,111,109]
-
Delphine Serpin
Concepto Azul S.A.
P.O. Box 0902-142A
Guayaquil, Ecuador -
Tayron Vera
Concepto Azul S.A.
P. O. Box 0902-142A
Guayaquil, Ecuador -
Mayra Valdez
Concepto Azul S.A.
P.O. Box 0902-142A
Guayaquil, Ecuador -
Efraim Cayra
Marinazul S.A.
Lima, Peru
Tagged With
Related Posts

Health & Welfare
Aiding gut health with a natural growth promotor
A study with Nile tilapia conducted in commercial production cages in Brazil showed the potential – in the absence of major disease threats – of a commercial, natural growth promotor that modulates the microbiota (inhibiting growth of pathogenic bacteria and promoting growth of beneficial bacteria) and inhibits quorum sensing.
Innovation & Investment
Better together: Partnerships drive innovation at leading labs
Laboratories with industry partnerships are making aquaculture more innovative, efficient and responsible. These collaborations offer access to expertise, facilities and funding to further the industry and improve global food security.

Health & Welfare
Do current shrimp practices favor EMS?
After refilling ponds, surviving microorganisms – including Vibrio parahaemolyticus, which causes EMS in shrimp – may benefit from the availability of nutrients in sediment and water and lack of competing microorganisms.

Health & Welfare
Do probiotics work in aquaculture?
Probiotics administered in feeds provide competitive exclusion of pathogenic bacteria, create conditions unfavorable for pathogens and modulate intestinal immune responses.



